The scientific method & experimental design
Lecture 3
Thursday 26th March, 2026
Populations and samples
Populations and samples
Why we collect data
- Recording some kind of observation or measurement
- Example:
- Measuring the heights of different trees in a forest
- Measuring the carbon in the forest soil at different locations
- We want to say something about the forest in general

Photo by Olena Bohovyk
Populations and samples
Why we collect data
- Cannot measure every tree or soil at every location
- Instead, we collect a sample of data
- Use the sample to draw conclusions about the population
- Statistics allows us to approximate properties of entire populations from a limited number of samples1

Photo by Olena Bohovyk
Populations and samples
Definitions
Population
The totality of individual observations about which inferences are to be made, existing anywhere in the world or at least within a definitely specified sampling area limited in space and time.
Sample
A collection of individual observations selected by a specified procedure.
Populations and samples
Examples
Populations
- All the spruce (gran) trees in Skåne
- All the blue tits (blåmes) in Sweden
- All the genes in the common fruit fly (Drosophila melanogaster)
- All the herring (sill) in the Baltic sea
Samples
- 300 spruce trees from forests in Skåne
- 100 caught blue tits from nest boxes in Sweden
- 20 genes from the Drosophila melanogaster genome
- 1000 herring caught by a fishing boat off the coast of Karlskrona
Populations and samples
Parameters and statistics
- Many statistical analyses are focused on a numerical summary.
- E.g. Mean, standard deviation, correlation
- Can be exactly calculated (measurement error aside) from the population: population parameter
- Can be inferred from a representative sample: sample statistic
- If the data was collected representatively, then a sample statistic should be a good approximation of population parameter.
Populations and samples
Anecdotal evidence
“I saw a bumblebee in Skrylle that was huge! Therefore bumblebees in Skrylle must be unusually large.”
Populations and samples
Anecdotal evidence
“I saw a bumblebee in Skrylle that was huge! Therefore bumblebees in Skrylle must be unusually large.”
- Few data points
- Data collected haphazardly
- Rare cases are more memorable than common ones
Populations and samples
How to sample from a population
How could we collect a representative sample of:
- Students currently studying at the Department of Biology?
- Students across the whole university?

Photo by Alexandra Roslund
02:30
Populations and samples
How to sample from a population
How could we collect a representative sample of:
- DNA from red squirrels in Skåne?

02:30
Populations and samples
How to sample from a population
- If we want to claim that our sample statistic is a good representation of the population parameter:
- Sample is unbiased
- Randomness is a good way to achieve that
- But sometimes simple random sampling is not appropriate
Populations and samples
If you know the population parameter, no need for statistics
Experimental design
Experimental design
Observational vs experimental studies
What are the main difference between these two studies?
- I measure the biomass of wild Mercurialis annua plants found in sandy soils and in loamy soils in a nature reserve.
- I grow Mercurialis annua plants in either sandy or loamy soils from seeds, and measure there biomass after a 3 months.
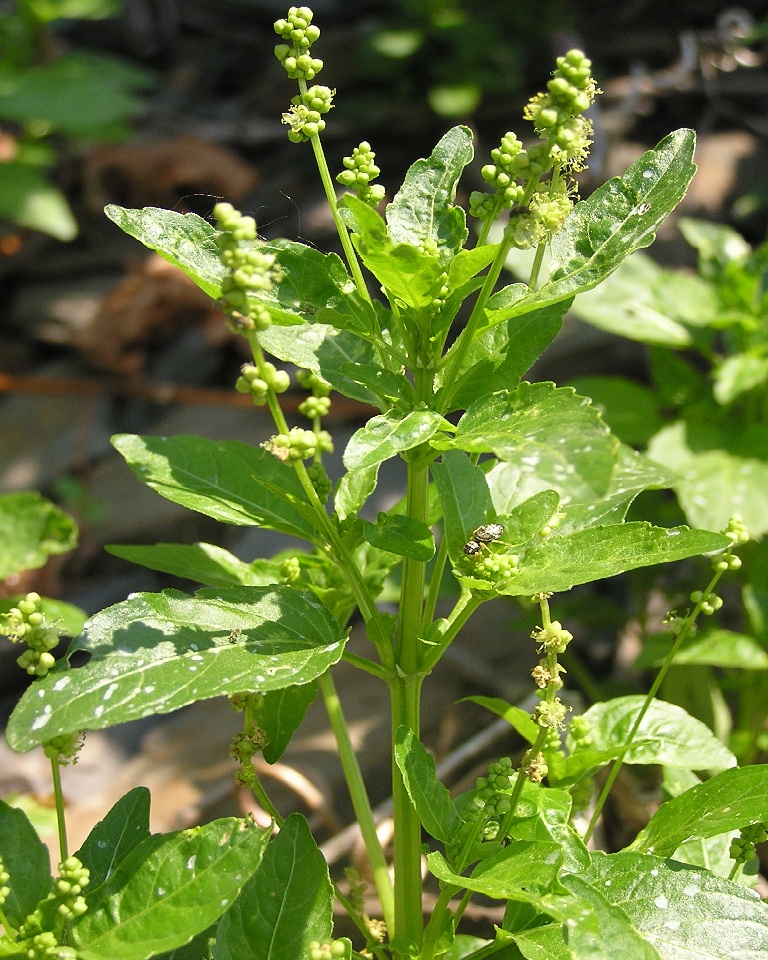

Photo by Michael Becker
02:00
Experimental design
Principles of experimental design
Experiment:
- When we assign treatments
- When we make an intervention
- When we manipulate something
- No longer just observing
Experimental design
Principles of experimental design: controlling
Try to control for differences that we can control but are not interested in.
For example:
- Water all plants the same amount
- Keep the temperature in the greenhouse the same
- Space out the plants evenly

Photo by Michael Becker
Experimental design
Principles of experimental design: randomisation
Try to account for differences that we cannot control and are not interested in.
For example:
- Randomly assign seeds to soil type (treatment)
- Randomly assign pots to rooms in a greenhouse

Photo by Michael Becker
Experimental design
Principles of experimental design: replication
Which statement gives you more confidence? Why?
“A clinical trial of a new blood pressure medication reduced the number of heart attacks in the treatment group by 96% and no negative side effects were reported (sample size = 14 people)”
“A clinical trial of a new blood pressure medication reduced the number of heart attacks in the treatment group by 82%, and 2% of participants reported negative side effects (sample size = 300 people)”
02:00
Experimental design
Principles of experimental design: replication
- The larger the sample size, the more accurately we can assess the effect of our treatment (explanatory variable) on the response variable.
- Each replicate should be independent of all others
- Otherwise we risk pseudoreplication
Experimental design
Principles of experimental design: replication
Pseudoreplication
- 50 plants in each treatment
- I measure 10 leaves from each plant
- Is my sample size per treatment:
- n = 50
- n = 500

Photo by Michael Becker
02:00
Experimental design
Principles of experimental design: blocking
When we suspect variables other than the treatment influence the treatment. Sometimes done for logistical reasons.
Examples:
- Temporal blocks: split into experimental groups that are conducted at different times
- Spatial blocks: split into experimental groups that are conducted in different locations
- “Risk” blocks: split into experimental groups that you expect to react differently to the treatment
Experimental design
Common types of experimental design: factorial design
Experiments where multiple treatments are applied, and all combinations of treatments are used:
| Soil type | Fertiliser |
|---|---|
| Sandy | None |
| Sandy | Added |
| Loamy | None |
| Loamy | Added |
Causation
Causation
Observational vs experimental studies
What are the main difference between these two studies?
- I measure the biomass of wild Mercurialis annua plants found in sandy soils and in loamy soils in a nature reserve.
- I grow Mercurialis annua plants in either sandy or loamy soils from seeds, and measure there biomass after a 3 months.

Photo by Michael Becker
Causation
Causal pathways
Causation
Confounding variables
- Anything that confuses you about the causation
- Can be ommitted variables
Causation
Confounding variables
- Anything that confuses you about the causation
- Can be ommitted variables
- Can also be measured
Causation
Causal reasoning via DAGs
- Directed acyclic graphs
- Causation flows along the arrows
- A causes Y
- B also causes Y
- Used to define causal relationships to then:
- Design experiments
- Design statistical methods
Causation
Causal reasoning via DAGs
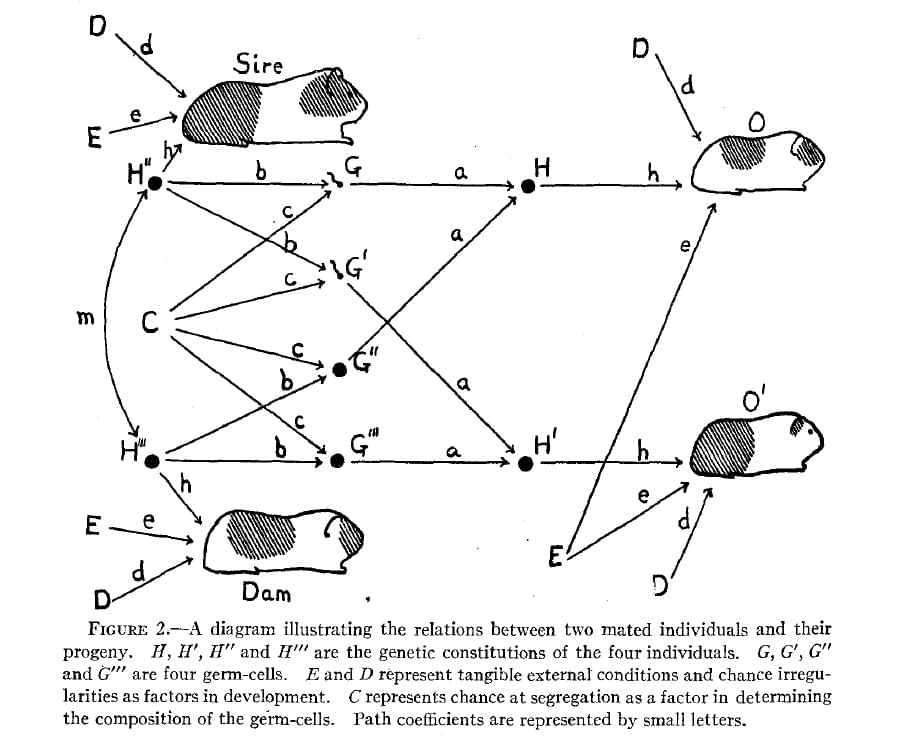
Causation
Causal reasoning via DAGs
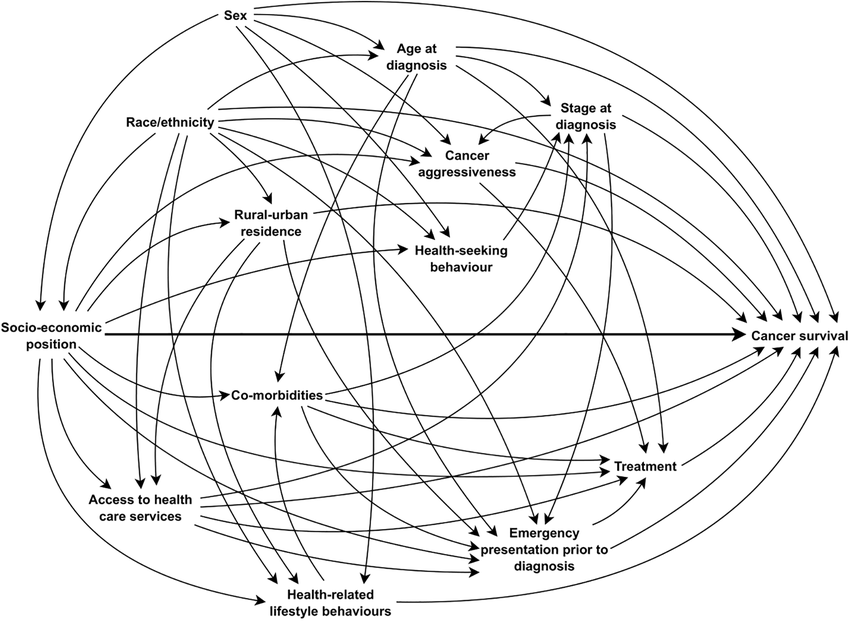
Causation
Causal reasoning via DAGs
Causation
Causal reasoning via DAGs: forks
- X and Y are associated (not independent)
- Z is a “common cause”
- Once grouped by Z, no association between X and Y
Causation
Causal reasoning via DAGs: forks
- X and Y are associated (not independent)
- Z is a “common cause”
- Once grouped by Z, no association between X and Y
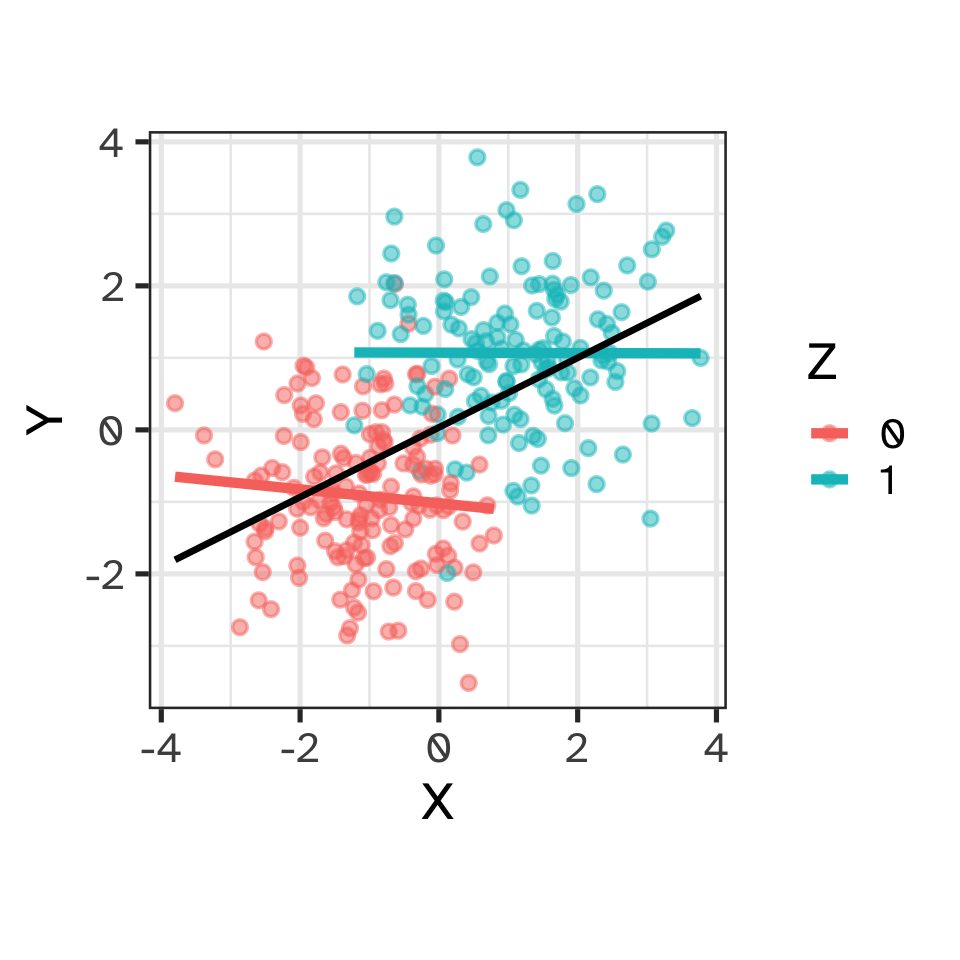
Causation
Causal reasoning via DAGs: pipes
- X and Y are associated (not independent)
- The effect of X on Y is transmitted through Z
- Once grouped by Z, no association between X and Y
Causation
Causal reasoning via DAGs: pipes
- X and Y are associated (not independent)
- The effect of X on Y is transmitted through Z
- Once grouped by Z, no association between X and Y
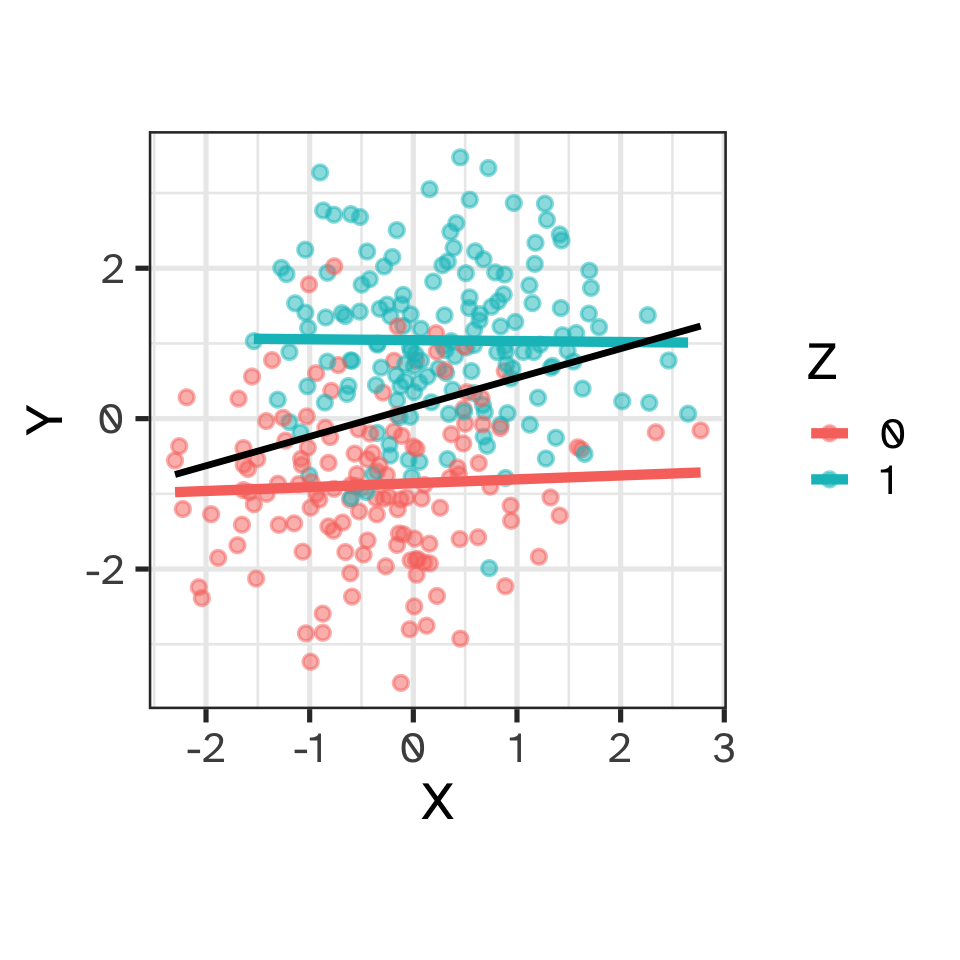
Causation
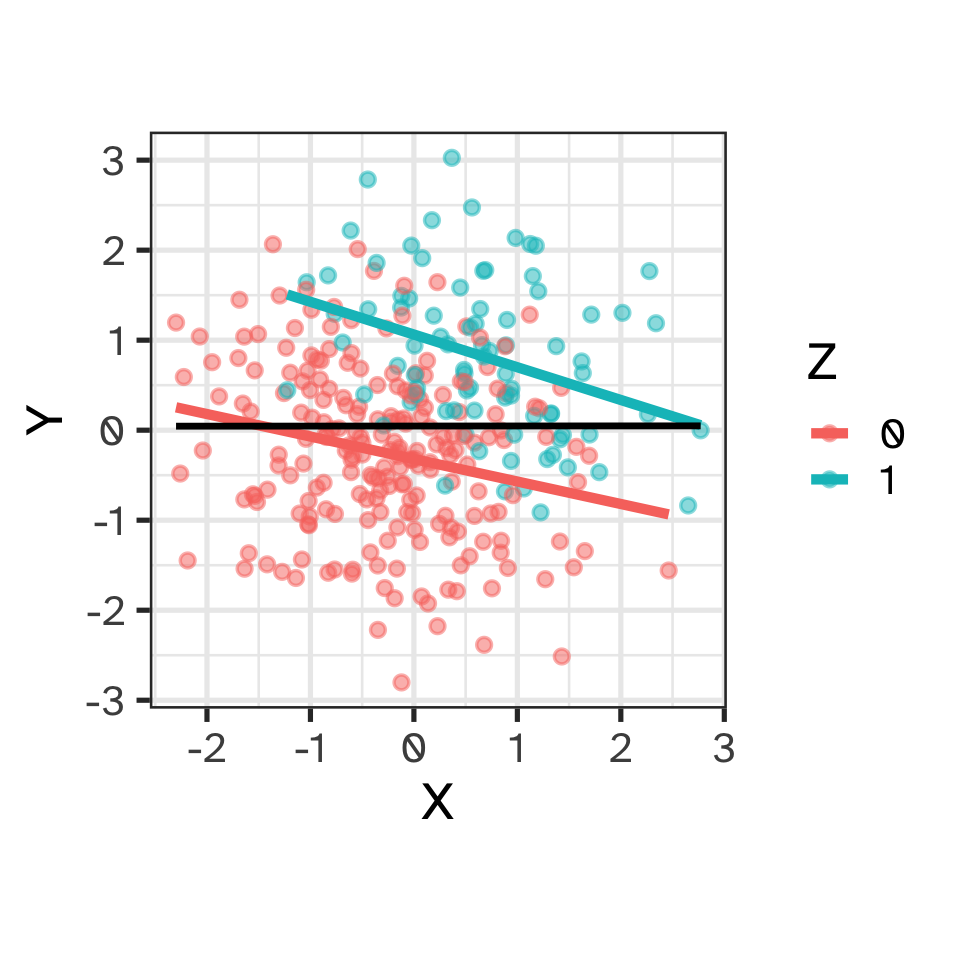
Causal reasoning via DAGs: colliders
- X and Y are not associated (independent)
- X and Y both influence Z
- Once grouped by Z, X and Y are associated
Causation
Causal reasoning via DAGs: colliders
- X and Y are not associated (independent)
- X and Y both influence Z
- Once grouped by Z, X and Y are associated
Causation
Causal reasoning via DAGs: descendants
- X and Y are causally associated via Z
- A contains information about Z
- Once grouped by A, X and Y are less associated
- A is a proxy for Z
The scientific method
The scientific method
Why do we do science?
- Why do you do science?
- Why should we (as a society) do science?
- Who do we do science for (if anyone)?
08:00
The scientific method
How do we do science?
- Why do we study what we study?
- Who decides?
- How do we find agreement?
- How do we handle disagreement?
- How do we go from unknowns to knowns?
- How does a scientific field progress?
12:00