Regression modelling
Lecture 6
Wednesday 1st April, 2026
Correlation
Correlation
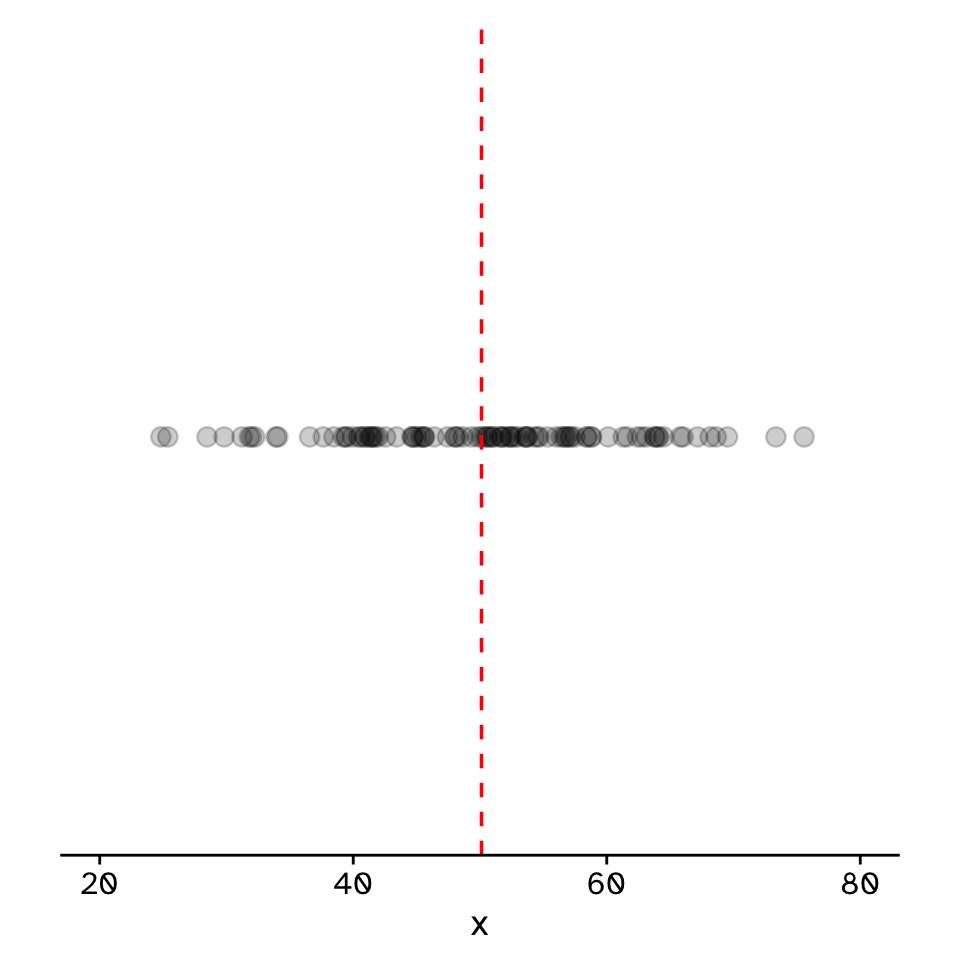
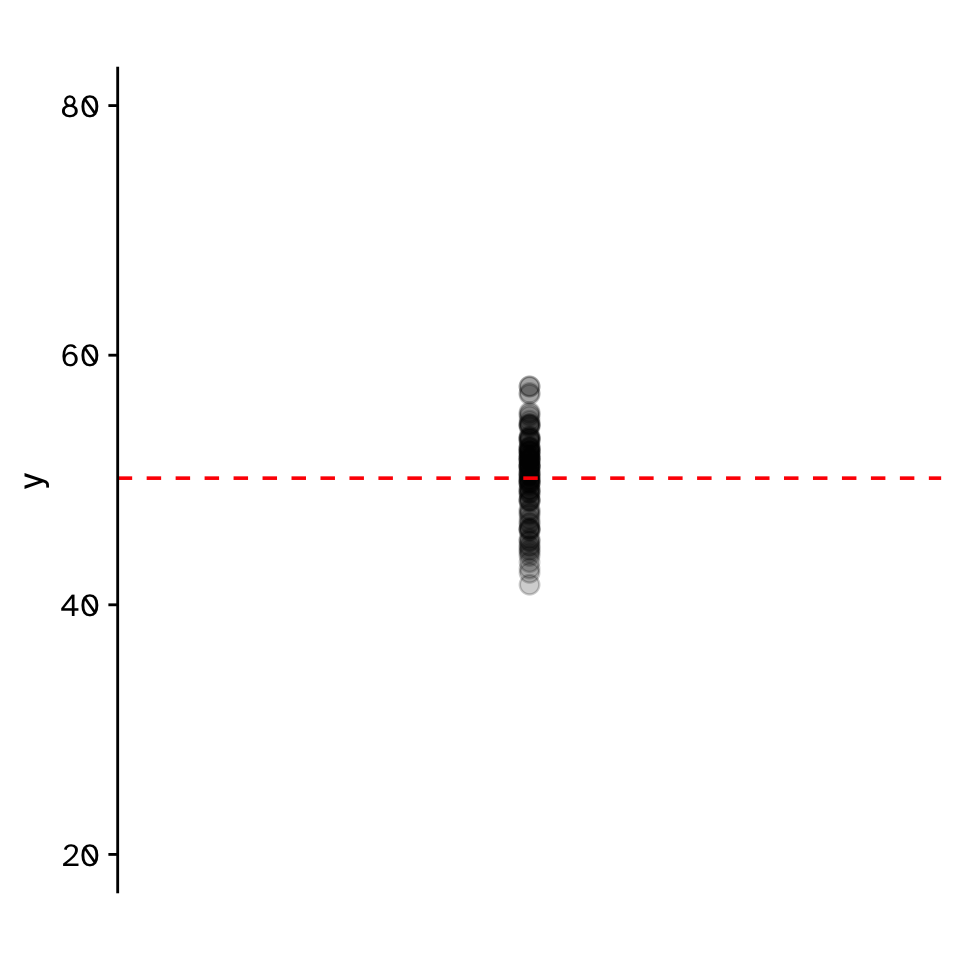
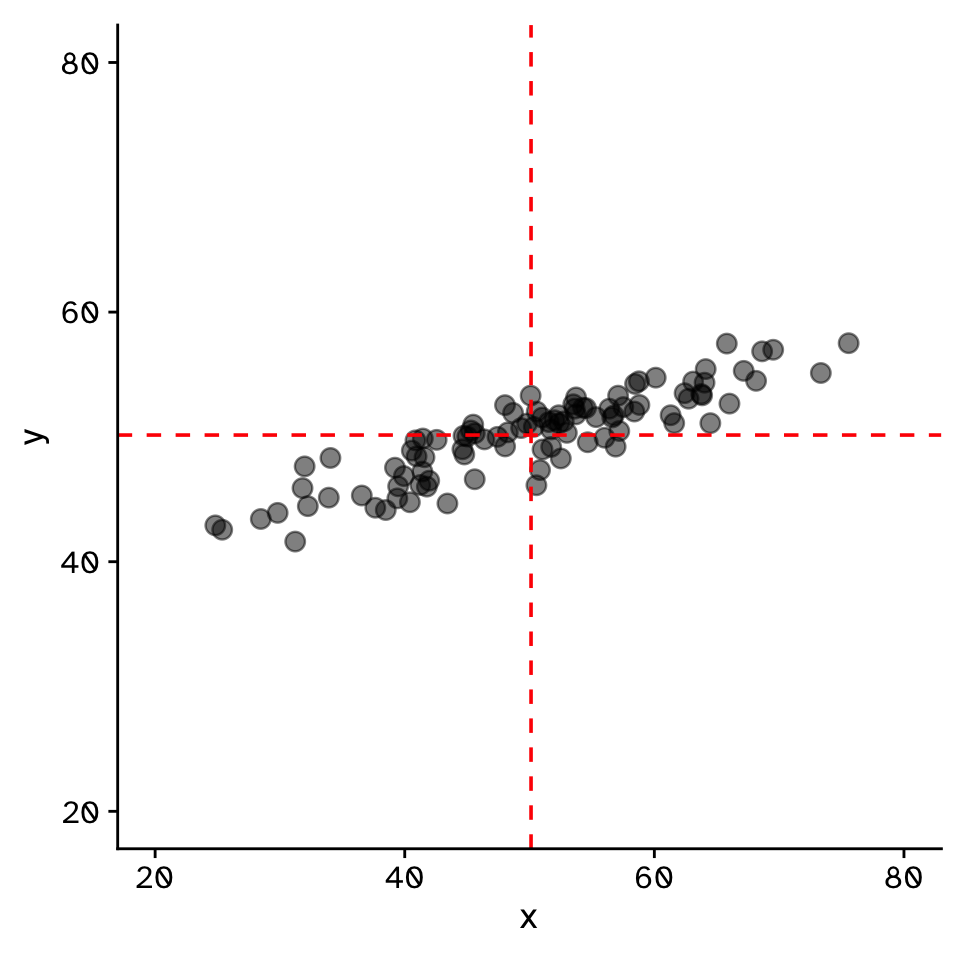
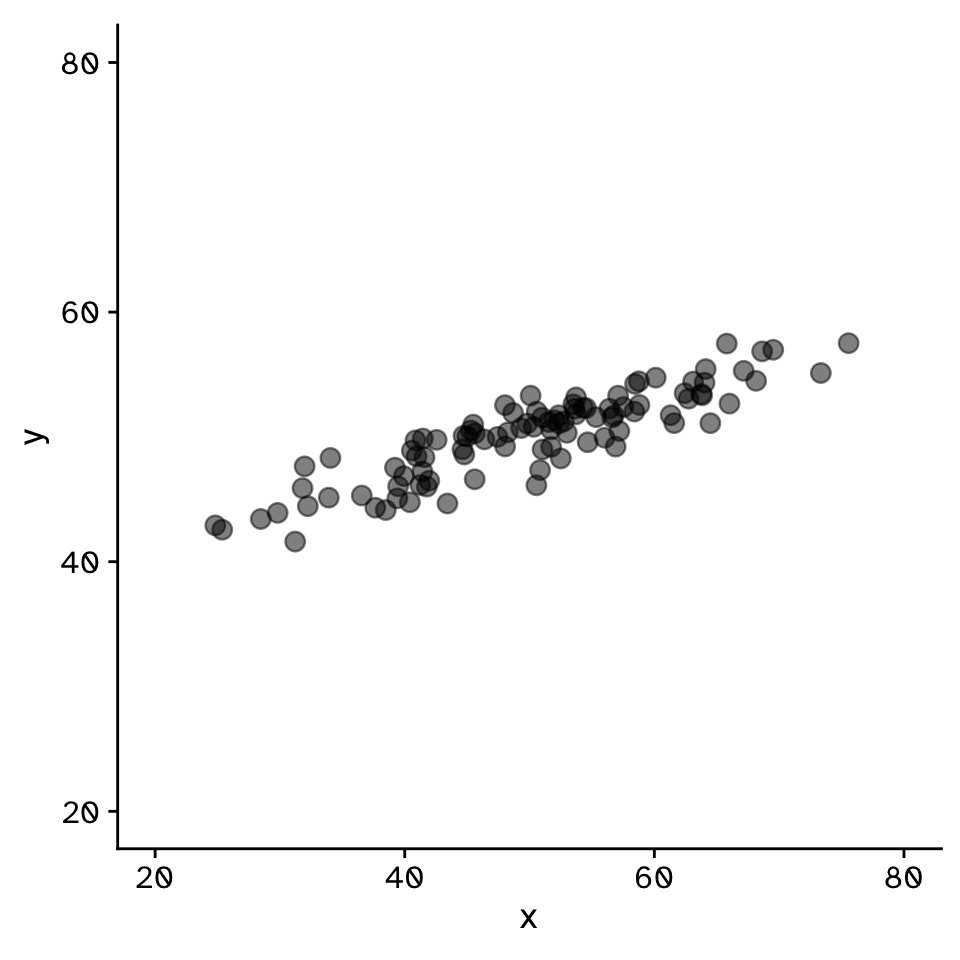
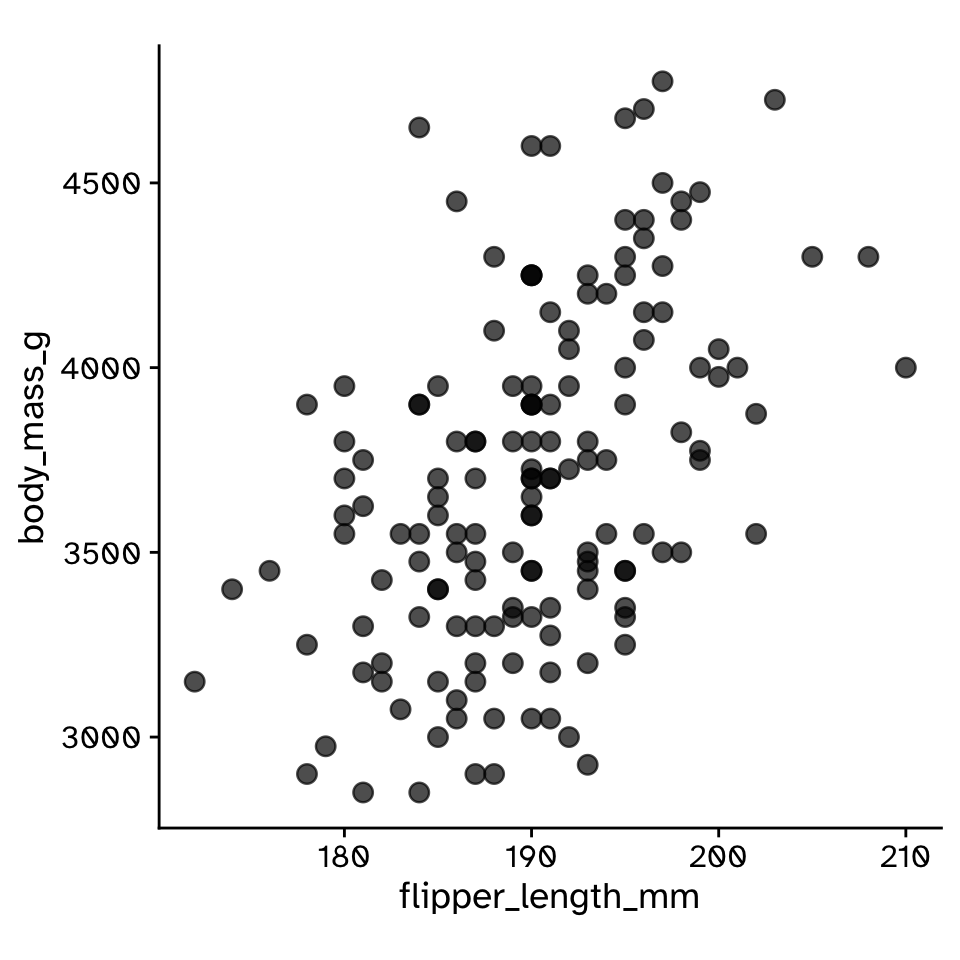
Do two continuous variables covary?
- Used to assess if two continuous variables are independent, or if they covary (linearly)
- We do not express one variable as a function of another:
- No “response” and “explanatory”
- Usually assumed that both variables are caused by the same thing
- “Correlation does not mean causation”
- But you can measure a correlation between a cause and effect
Correlation
Do two continuous variables covary?
Correlation coefficient (Pearson):
\[ r = \frac{\sum_{i=1}^n (x_i - \bar{x})(y_i - \bar{y})}{\sqrt{\sum_{i=1}^n (x_i - \bar{x})^2} \sqrt{\sum_{i=1}^n (y_i - \bar{y})^2}} \]
\[ r = \frac{\text{Covariance}(x,y)}{\text{Standard deviation}(x) \times \text{Standard deviation}(y)} \]
\[ r = \frac{\text{Cov}(x,y)}{\sigma_x \sigma_y} \]
Correlation
Do two continuous variables covary?
\[ r = \frac{\text{Cov}(x,y)}{\sigma_x \sigma_y} \]
Correlation
Do two continuous variables covary?
Correlation
Do two continuous variables covary?
Correlation
Do two continuous variables covary?
Correlation
Do two continuous variables covary?
Correlation
Do two continuous variables covary?
Correlation
Do two continuous variables covary?
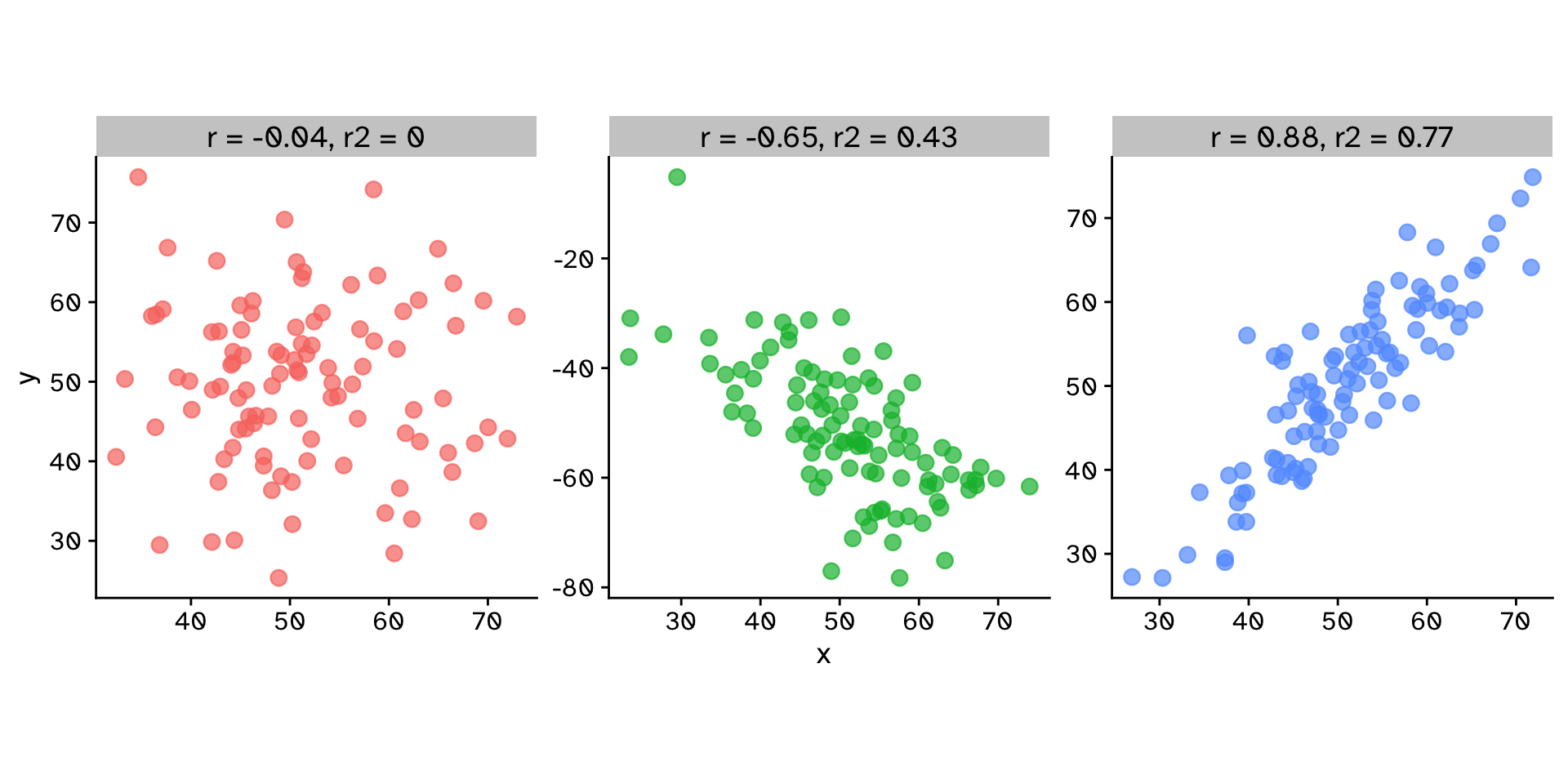
Correlation coefficient (\(r\)): \[ r = \frac{\sum_{i=1}^n (x_i - \bar{x})(y_i - \bar{y})}{\sqrt{\sum_{i=1}^n (x_i - \bar{x})^2} \sqrt{\sum_{i=1}^n (y_i - \bar{y})^2}} \]
Coefficient of determination (\(r^2\)): \[ r^2 = \left(\frac{\sum_{i=1}^n (x_i - \bar{x})(y_i - \bar{y})}{\sqrt{\sum_{i=1}^n (x_i - \bar{x})^2} \sqrt{\sum_{i=1}^n (y_i - \bar{y})^2}}\right)^2 \]
Correlation
Do two continuous variables covary?
Correlation
Do two continuous variables covary?
Correlation
Observed statistic
Correlation
Confidence intervals
Correlation
Confidence intervals
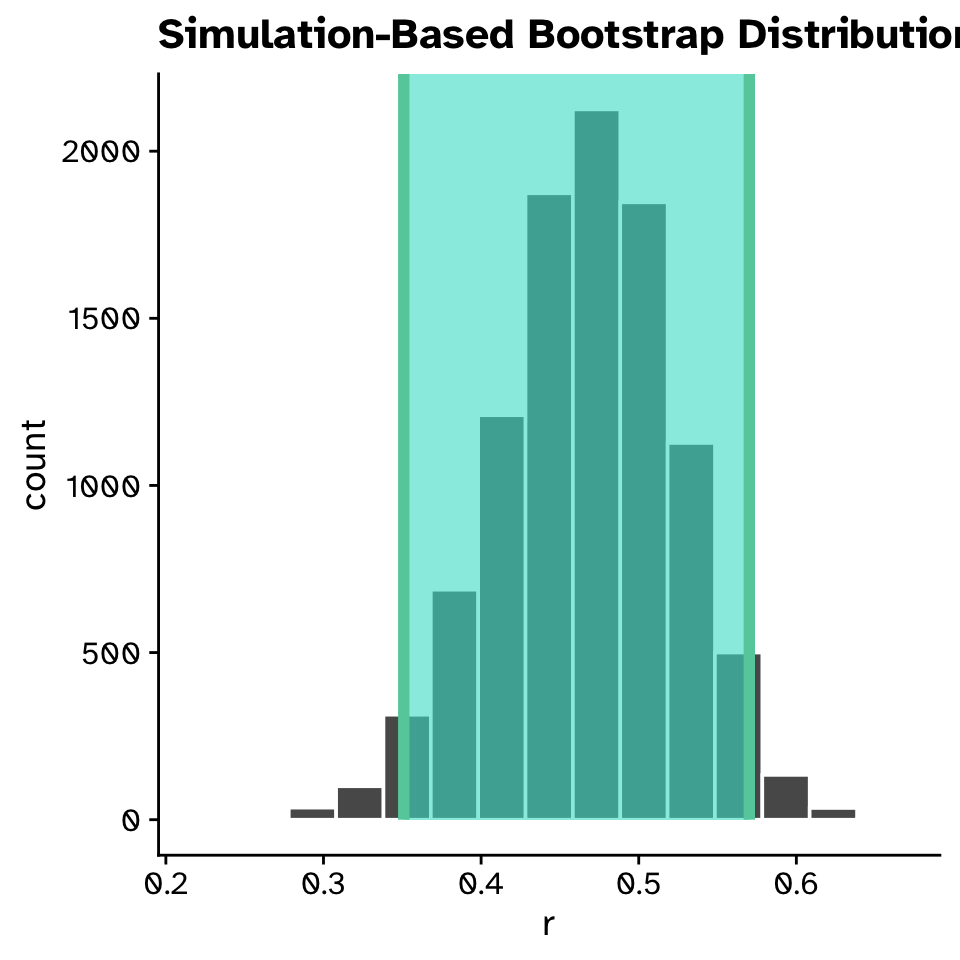
Correlation
Confidence intervals
- The correlation between flipper length and body mass was 0.468 (95% CI: 0.355, 0.572).
Correlation
Hypothesis test
- Null hypothesis:
- The two variables do not covary (\(r=0\))
- Alternative hypothesis:
- The two variables do covary (\(r\neq0\))
Correlation
Hypothesis test
- How could we generate a null distribution?
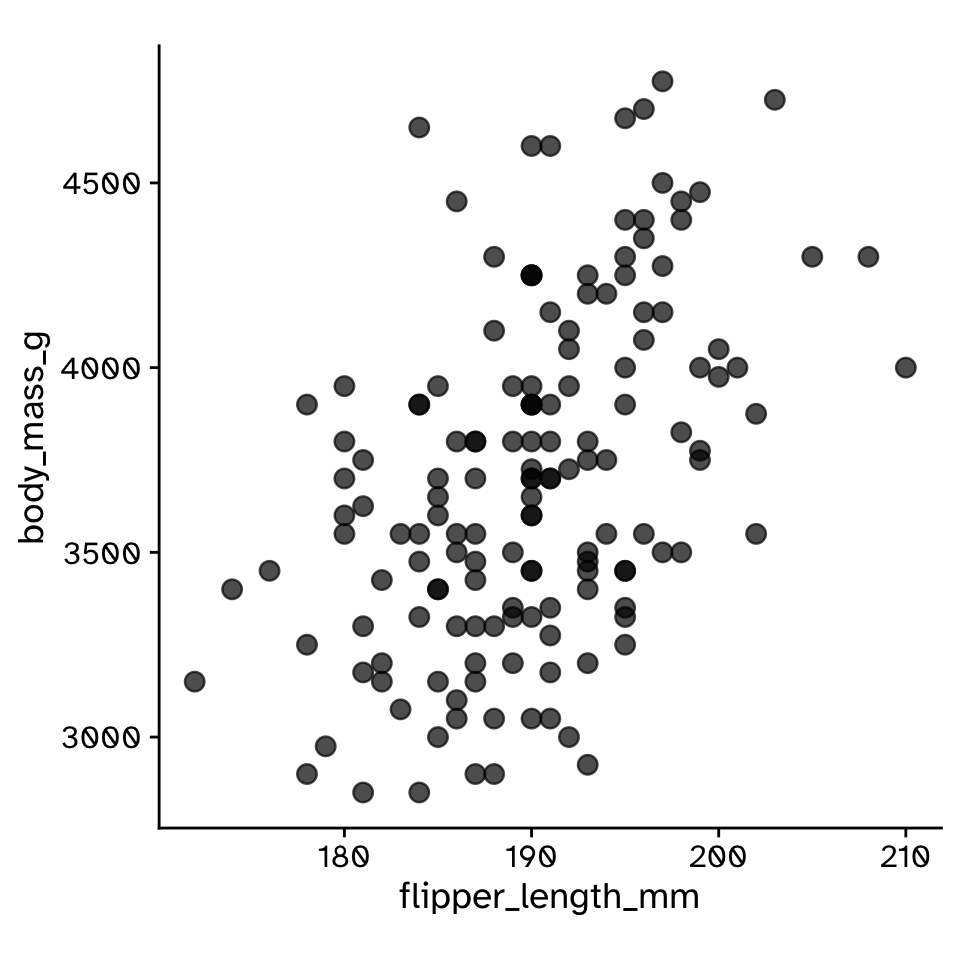
Correlation
Hypothesis test
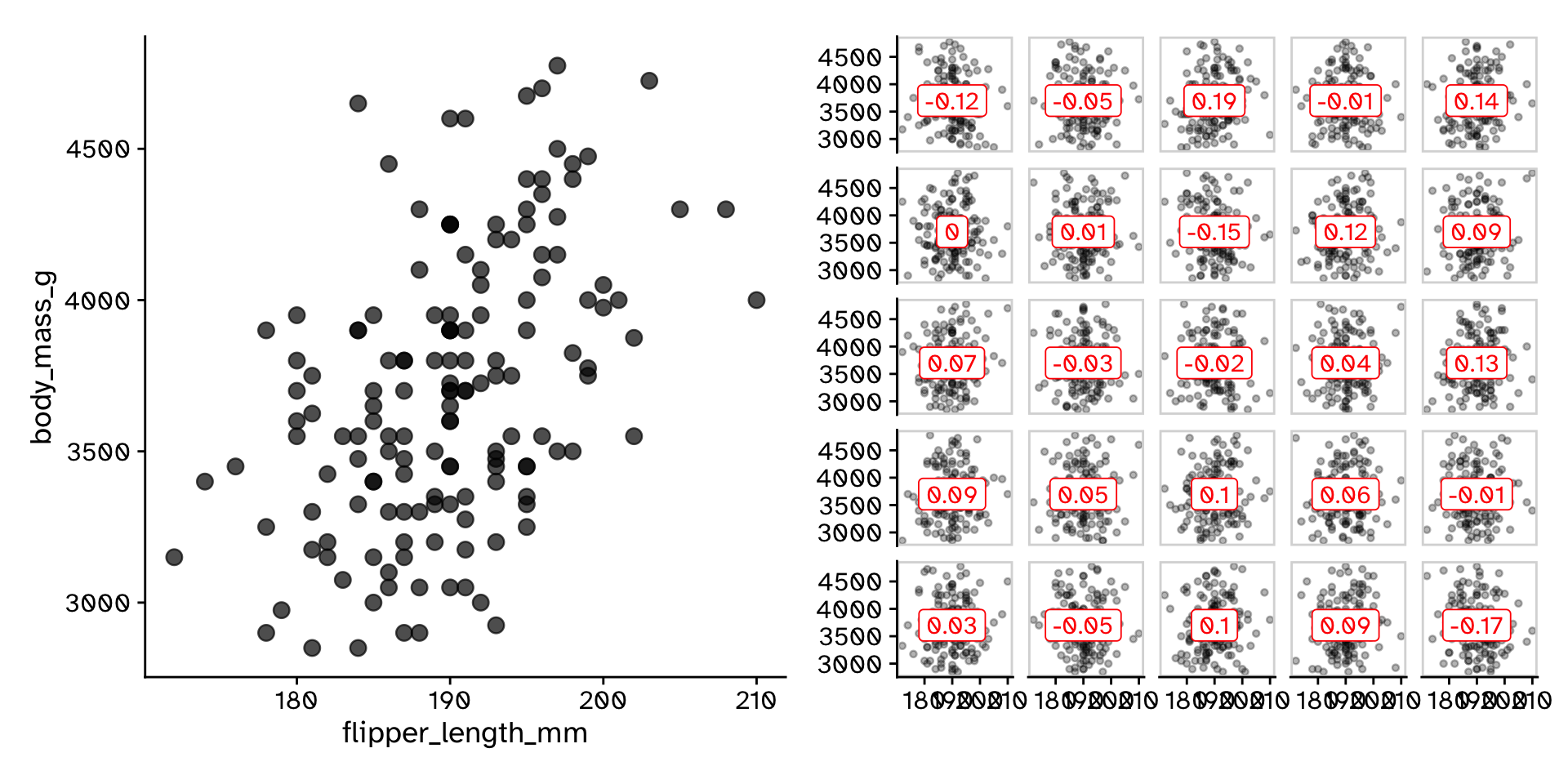
Correlation
Hypothesis test
Correlation
Hypothesis test
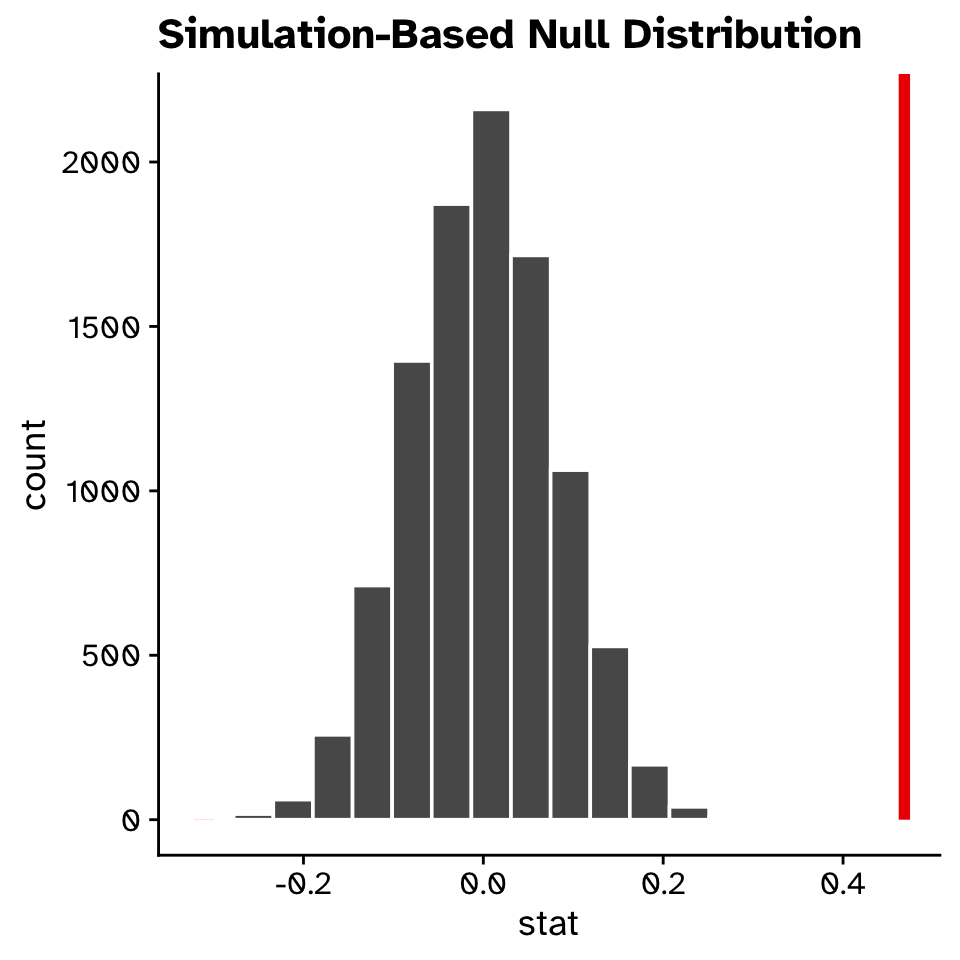
Correlation
Hypothesis test
# A tibble: 1 × 1
p_value
<dbl>
1 0- Remember that our p-value is an approximation
- The number of decimal places of accuracy we can observe is directly linked to the number of
reps
- The number of decimal places of accuracy we can observe is directly linked to the number of
Linear Regression
- When your response is a quantitative variable, and you want to predict it using (at least) one other quantitative variable
Linear Regression
How does variable Y depend on variable X
- A mathematical function describing the relationship between two continuous variables
- Y is dependant on X (the)
- Y is a function of X
- Predictive model
- Very powerful framework
Linear Regression
How does variable Y depend on variable X
\[ y = \text{Slope}\times x + \text{Intercept} \]
\[ y = mx+c \]
\[ y = \beta_1x+\beta_0 \]
\[ y = \beta_0+\beta_1x+\beta_2x_2+\beta_3x_3+\beta_4x_4+\beta_5x_5 \]
Linear Regression
How does variable Y depend on variable X
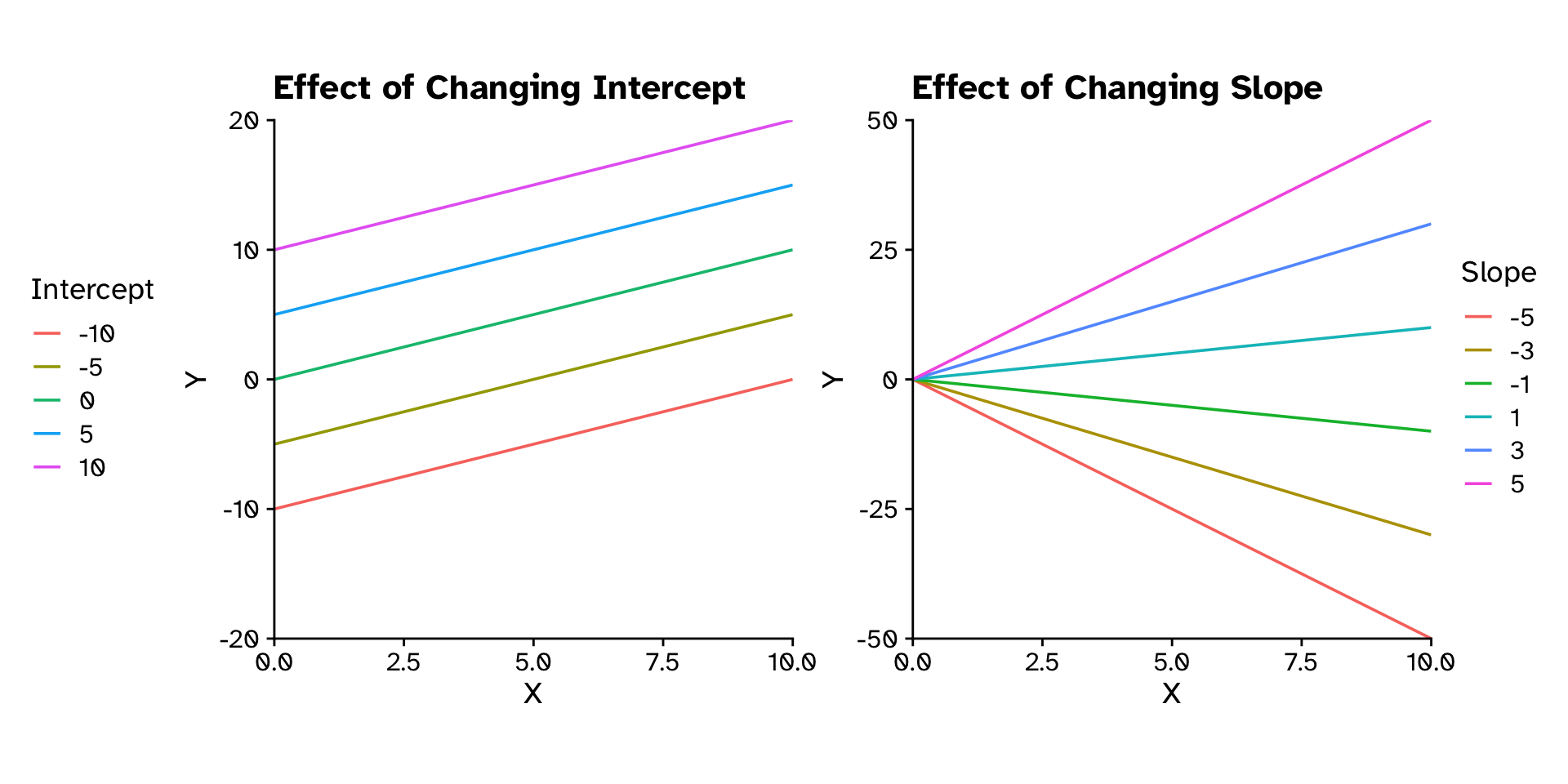
Linear Regression
How does variable Y depend on variable X
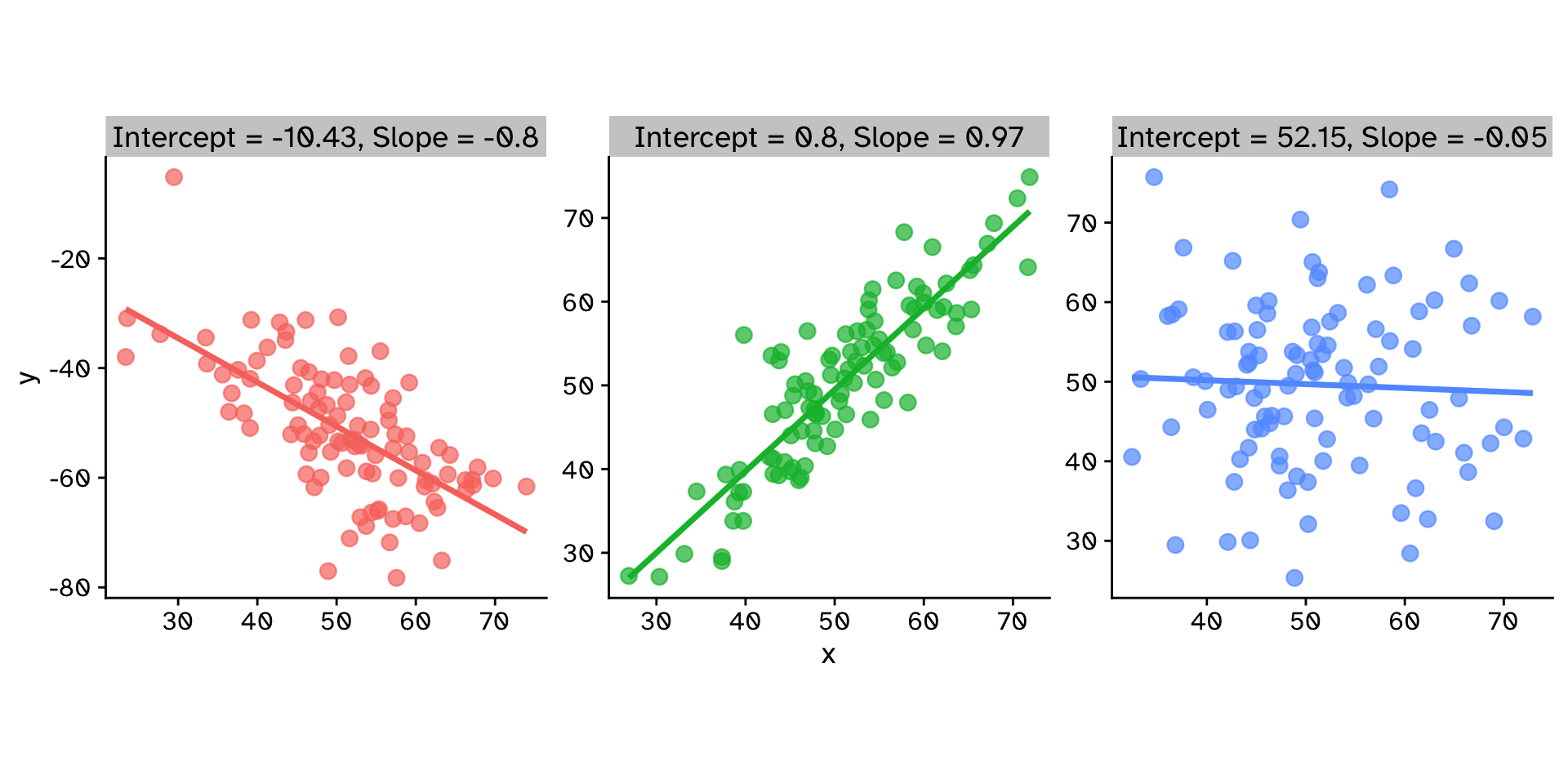
Linear Regression
How does variable Y depend on variable X
- Example: For sexual selection to operate, an increase in mating success (number of mates) must result in an increase in reproductive success (number of offspring).
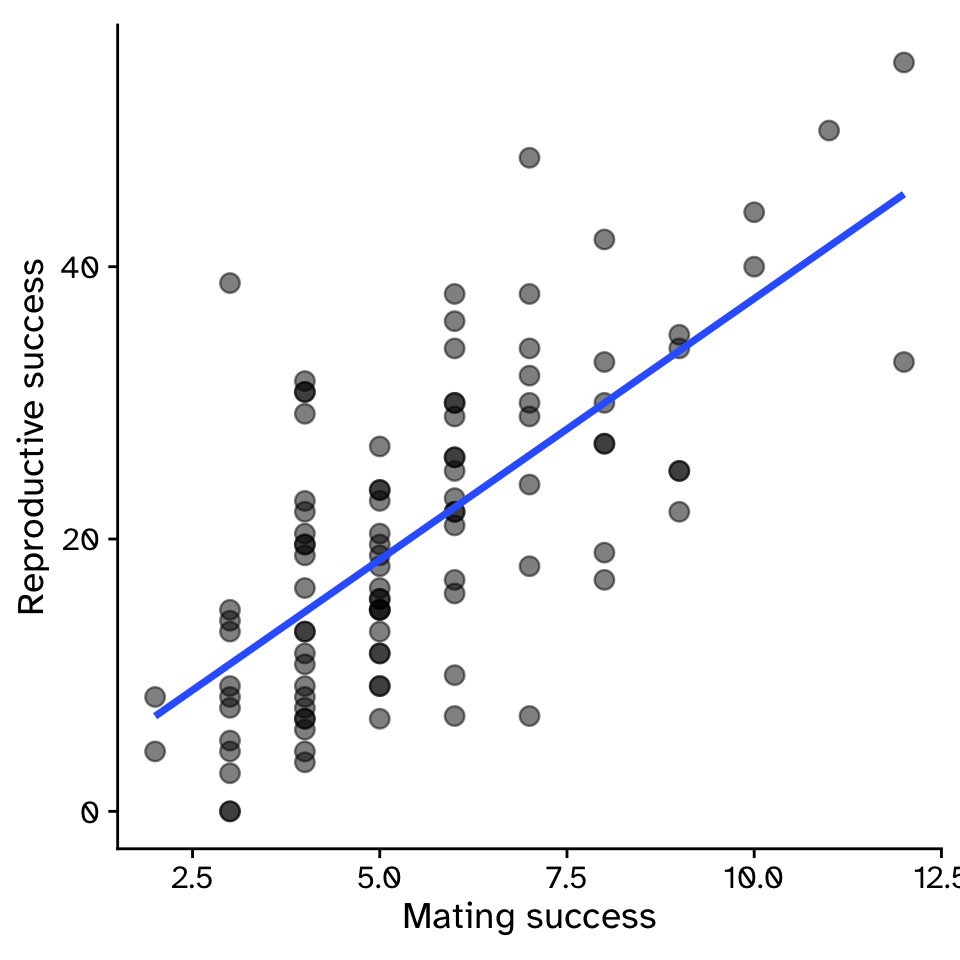
Linear Regression
How does variable Y depend on variable X
\[ y = \beta_1x+\beta_0 \]
\[ y = 3.83x-0.68 \]
- \(\beta_1\) = strength of sexual selection
- For each additional mate, an individual (on average) gains \(\beta_1\) additional offspring
- For 5 mates (\(x=5\)):
- \(y = 3.83\times5-0.68\)
- \(y = 18.47\)
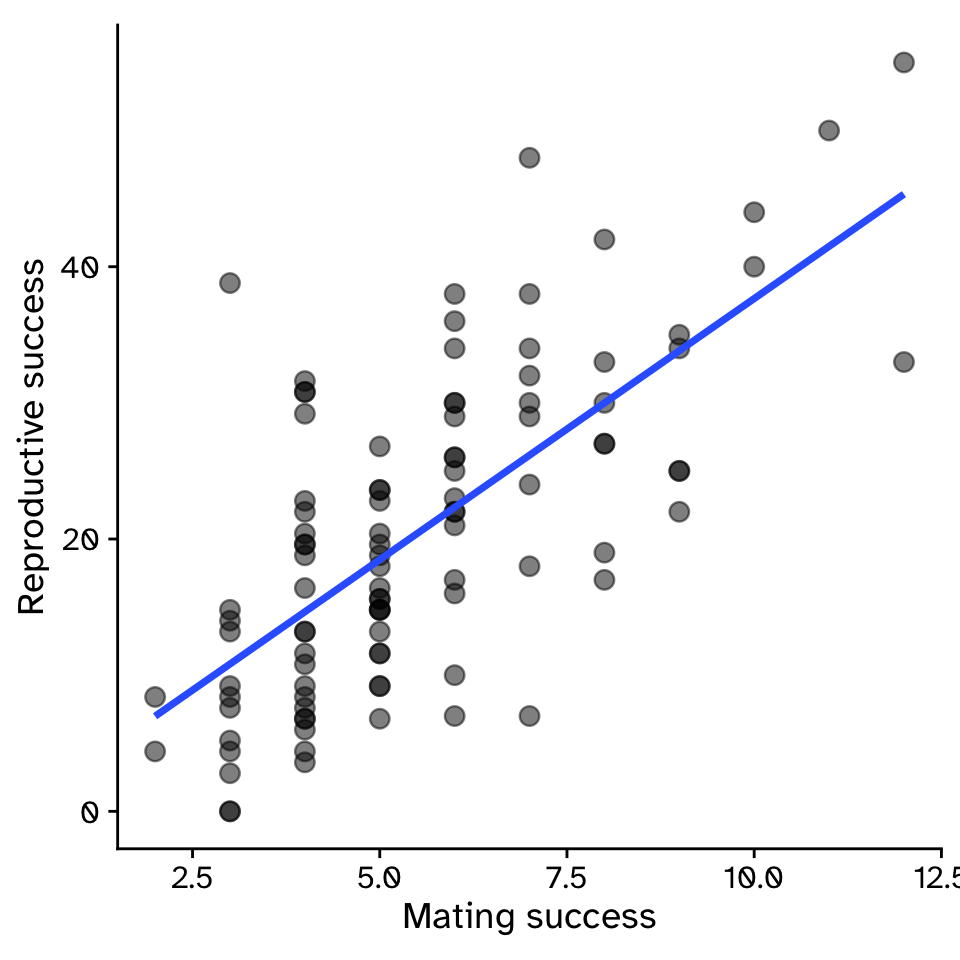
Linear Regression
How does variable Y depend on variable X
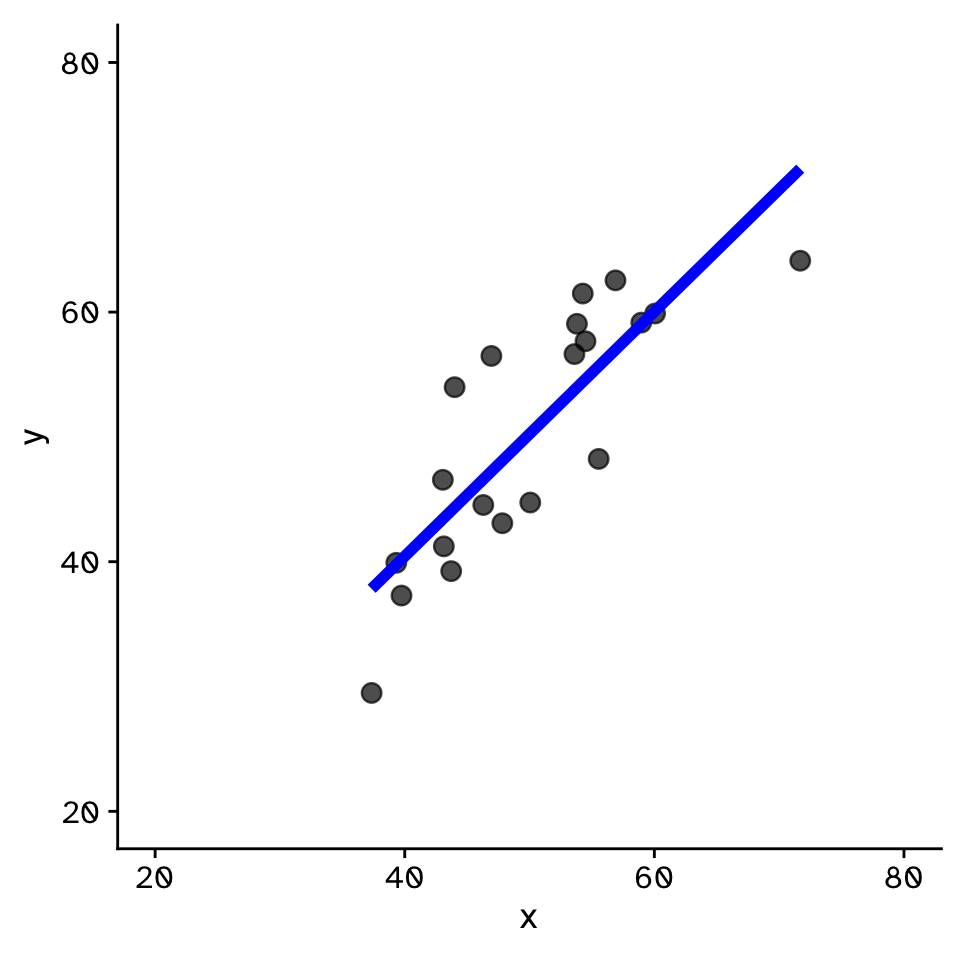
Linear Regression
How does variable Y depend on variable X
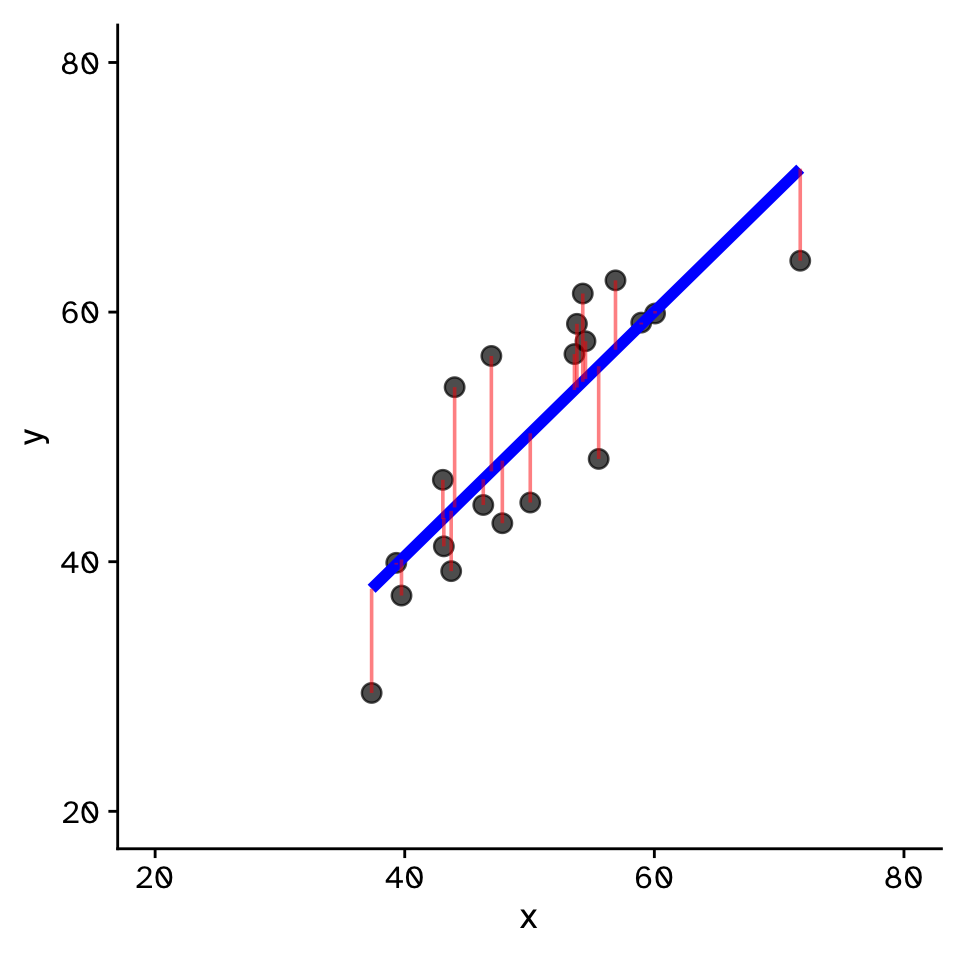
Linear Regression
How does variable Y depend on variable X
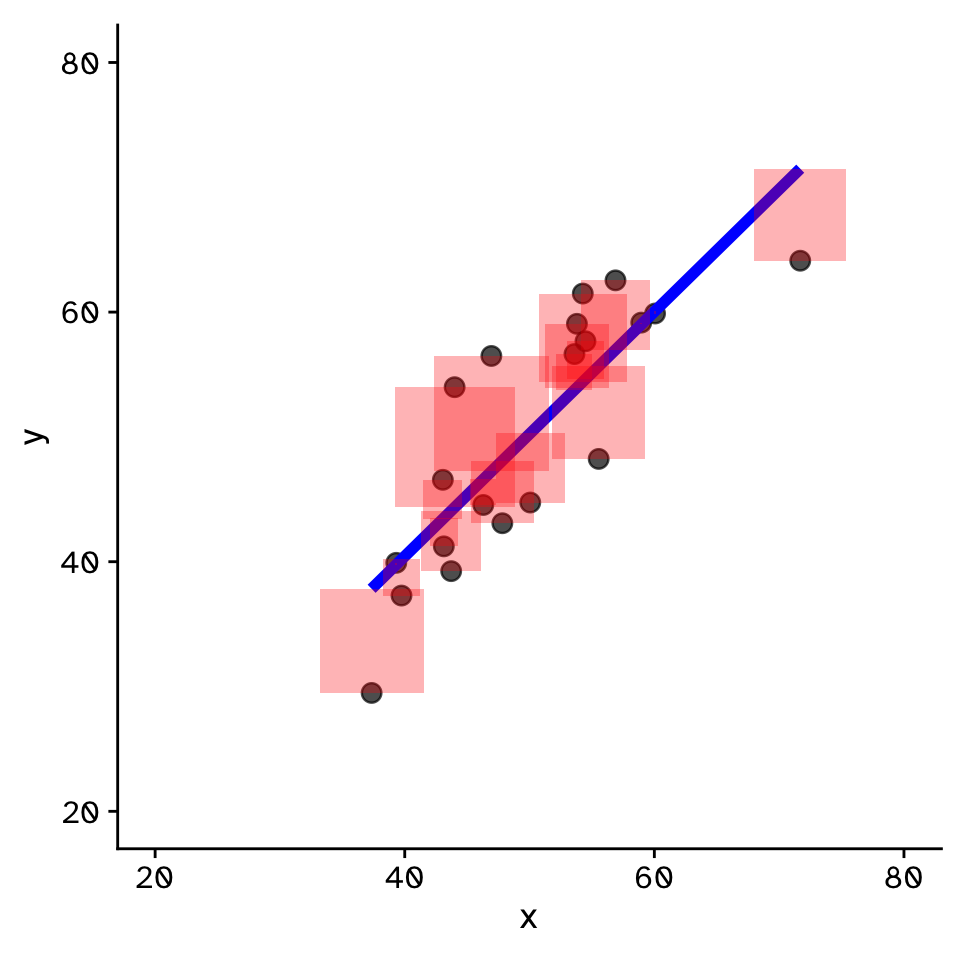
Linear Regression
How does variable Y depend on variable X
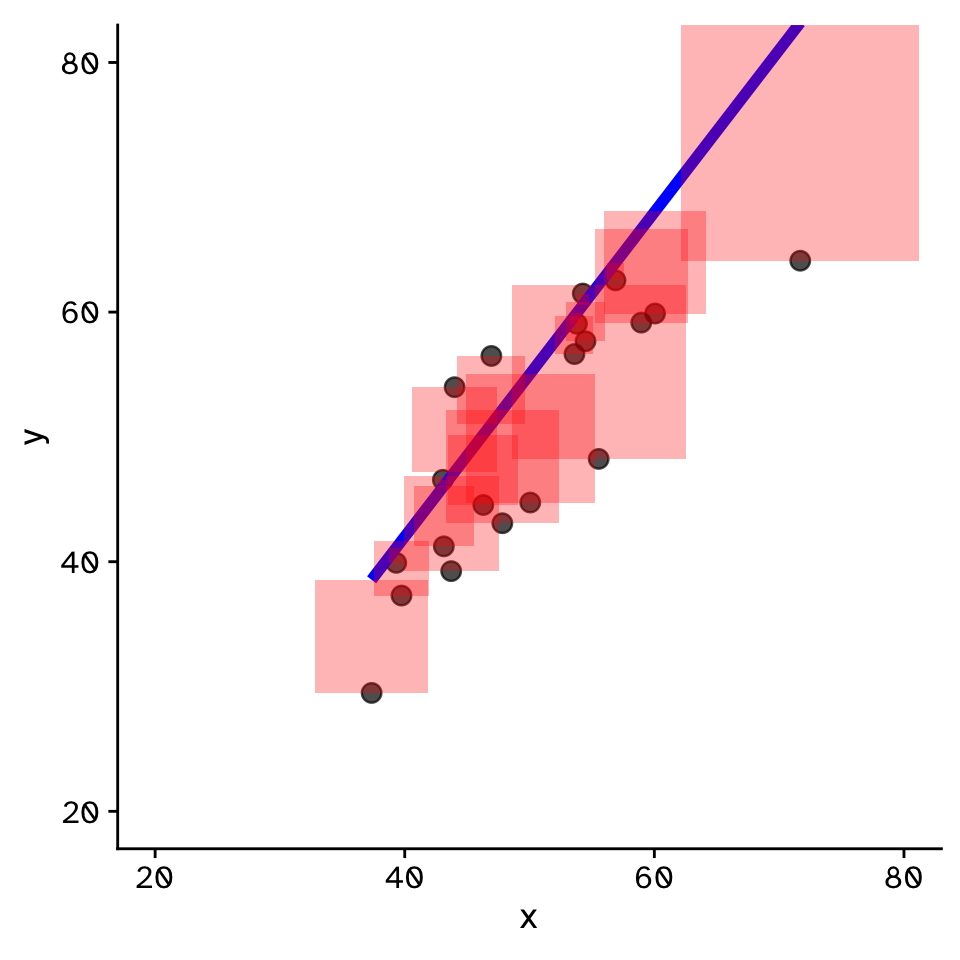
Linear Regression
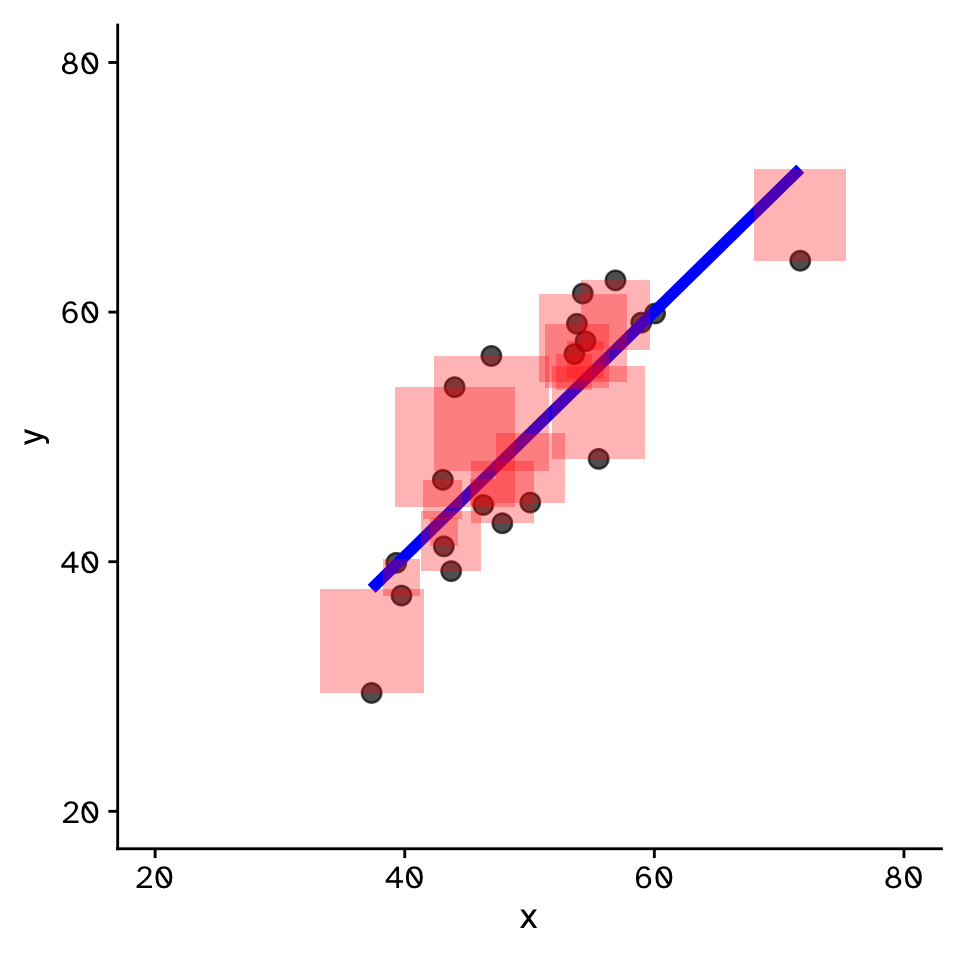
- Fit by solving to minimise the sum of the squared residuals (SSR)
- Find \(\beta_1\) and \(\beta_0\) that minimise the SSR
- Called a “loss function”
Linear Regression
How does variable Y depend on variable X
Linear Regression
How to fit a slope in R?
With infer (if no interest in the intercept):
Linear Regression
Confidence intervals for a slope
Response: reproductive_success (numeric)
Explanatory: mating_success (numeric)
# A tibble: 1,000 × 2
replicate stat
<int> <dbl>
1 1 4.32
2 2 4.73
3 3 3.80
4 4 2.84
5 5 3.96
6 6 3.98
7 7 3.38
8 8 3.60
9 9 4.08
10 10 3.98
# ℹ 990 more rowsLinear Regression
Confidence intervals for a slope
Linear Regression
Confidence intervals for a slope
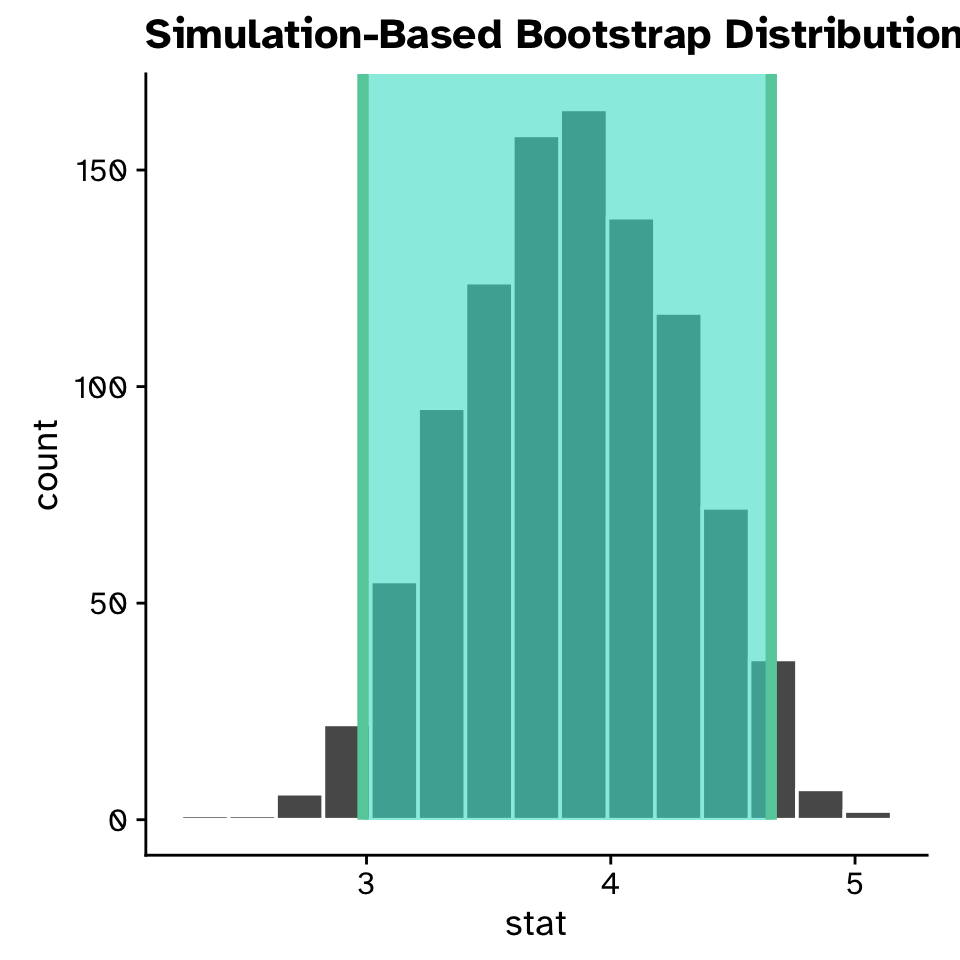
Linear Regression
Confidence intervals for a slope
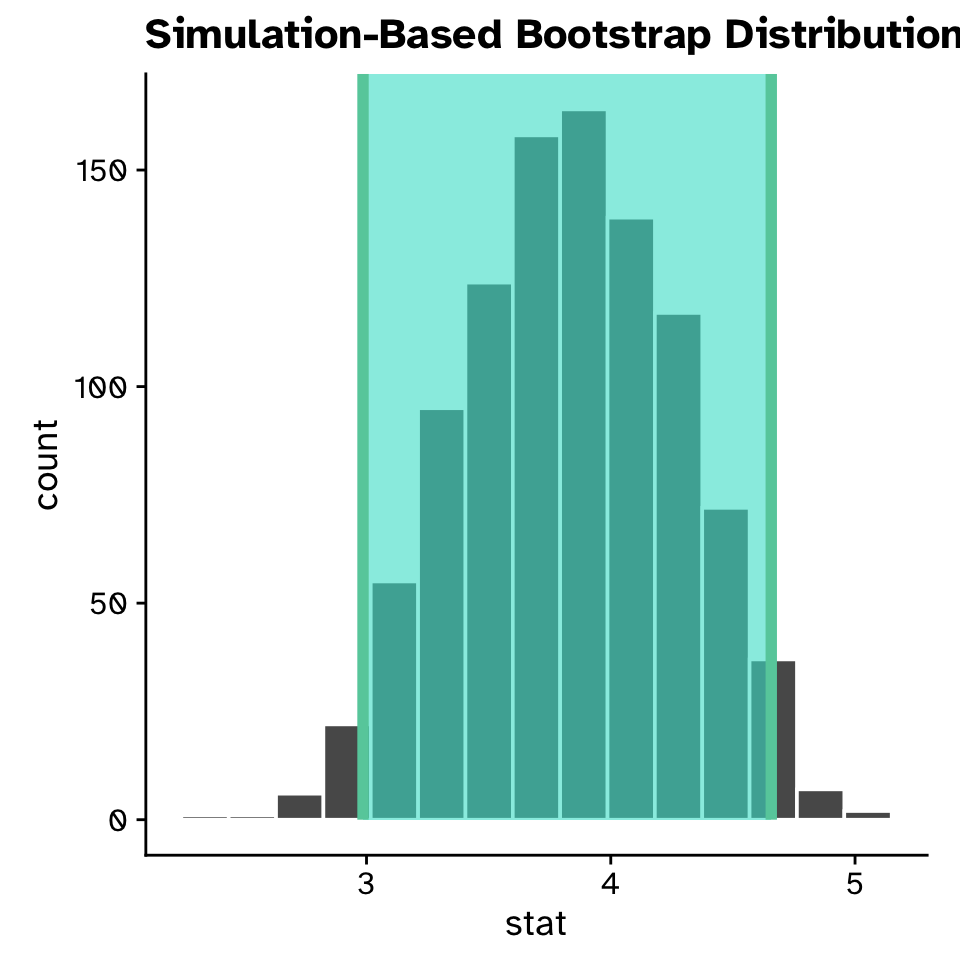
Linear Regression
Hypothesis test for a slope
- Null hypothesis:
- Slope = 0
- Alternative hypothesis:
- Slope \(\neq\) 0
- Slope < 0
- Slope > 0
Linear Regression
Hypothesis test for a slope
Response: reproductive_success (numeric)
Explanatory: mating_success (numeric...
# A tibble: 1,000 × 2
replicate stat
<int> <dbl>
1 1 0.0382
2 2 -0.603
3 3 -0.951
4 4 -0.570
5 5 -1.20
6 6 -0.0117
7 7 -0.548
8 8 -0.707
9 9 -0.423
10 10 -0.0407
# ℹ 990 more rowsLinear Regression
Hypothesis test for a slope
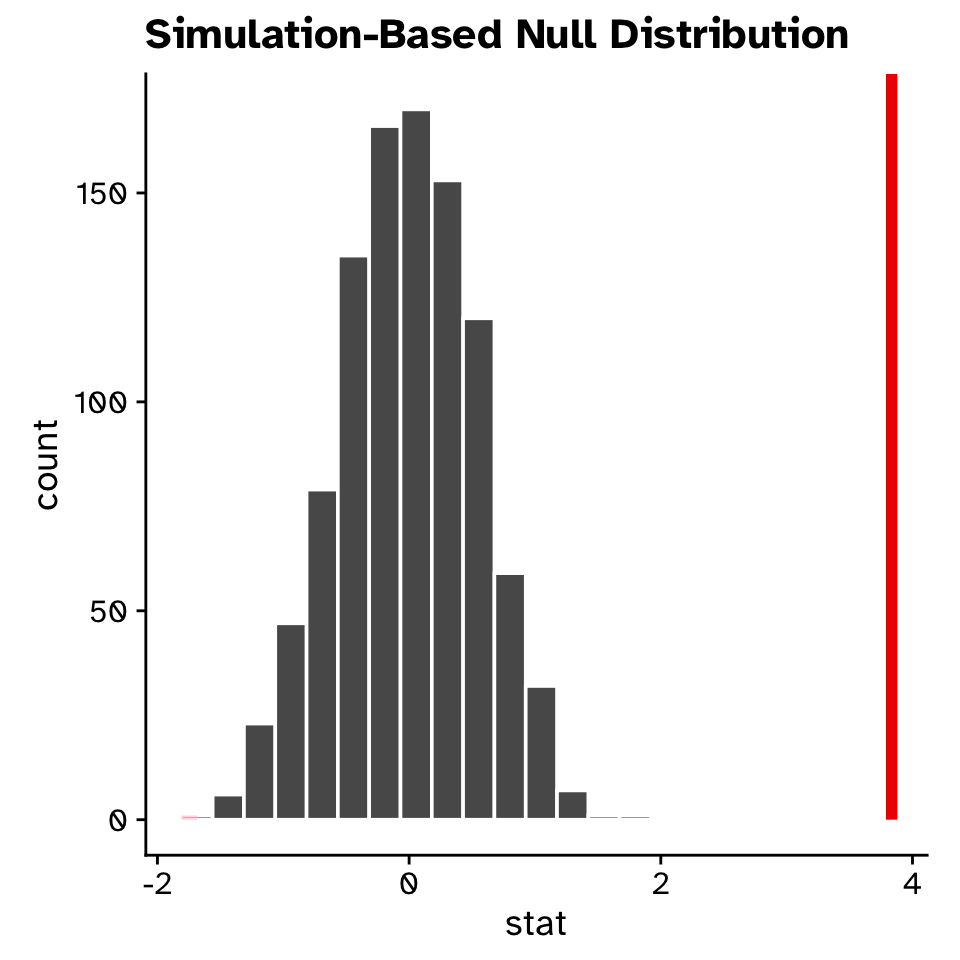
Linear Regression
Multiple linear regression
\[ y = \beta_1x+\beta_0 \]
\[ y = \beta_0+\beta_1x+\beta_2x_2+\beta_3x_3+\beta_4x_4+\beta_5x_5... \]
Linear Regression
Multiple linear regression: intercept only model
Linear Regression
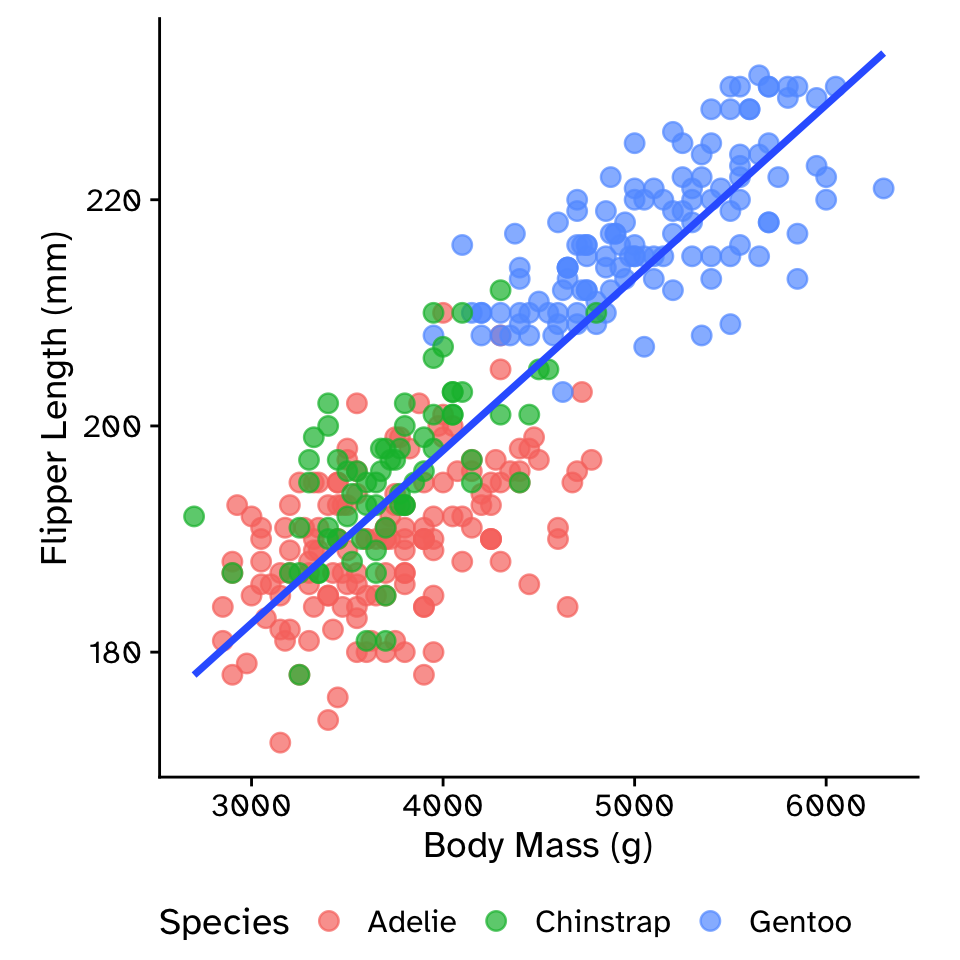
Multiple linear regression: intercept and slope model
Linear Regression
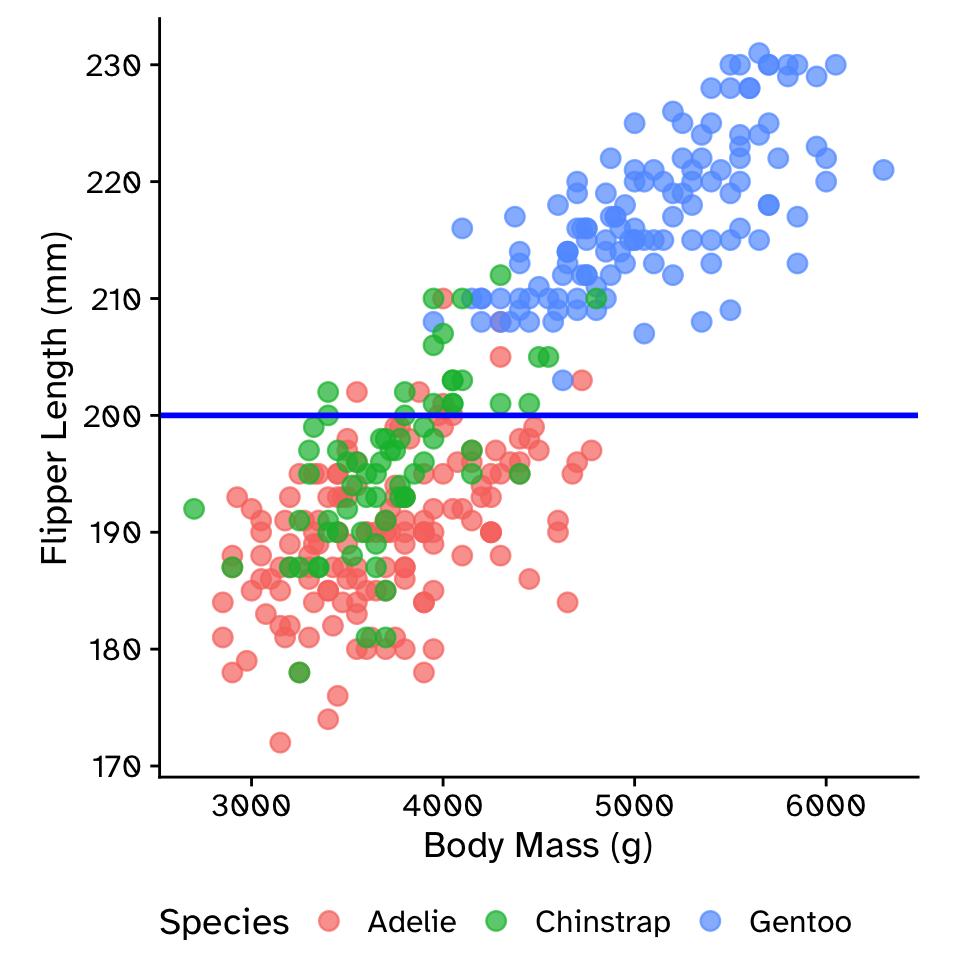
Multiple linear regression: additive model with parallel slopes (ANCOVA)
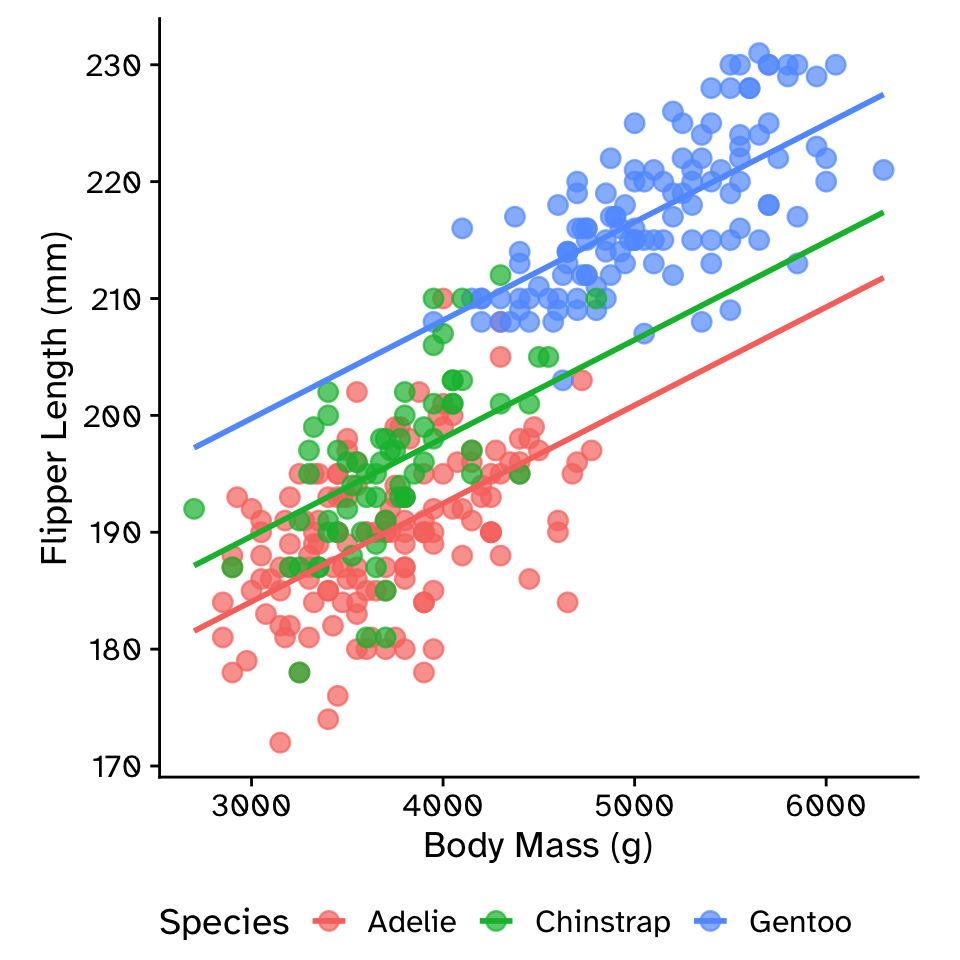
\[ \text{flipper_length_mm} = \beta_0+ \\ \beta_1 \times \text{body_mass_g}+ \\ \beta_2 \times \text{species} \]
Linear Regression
Multiple linear regression: interaction model
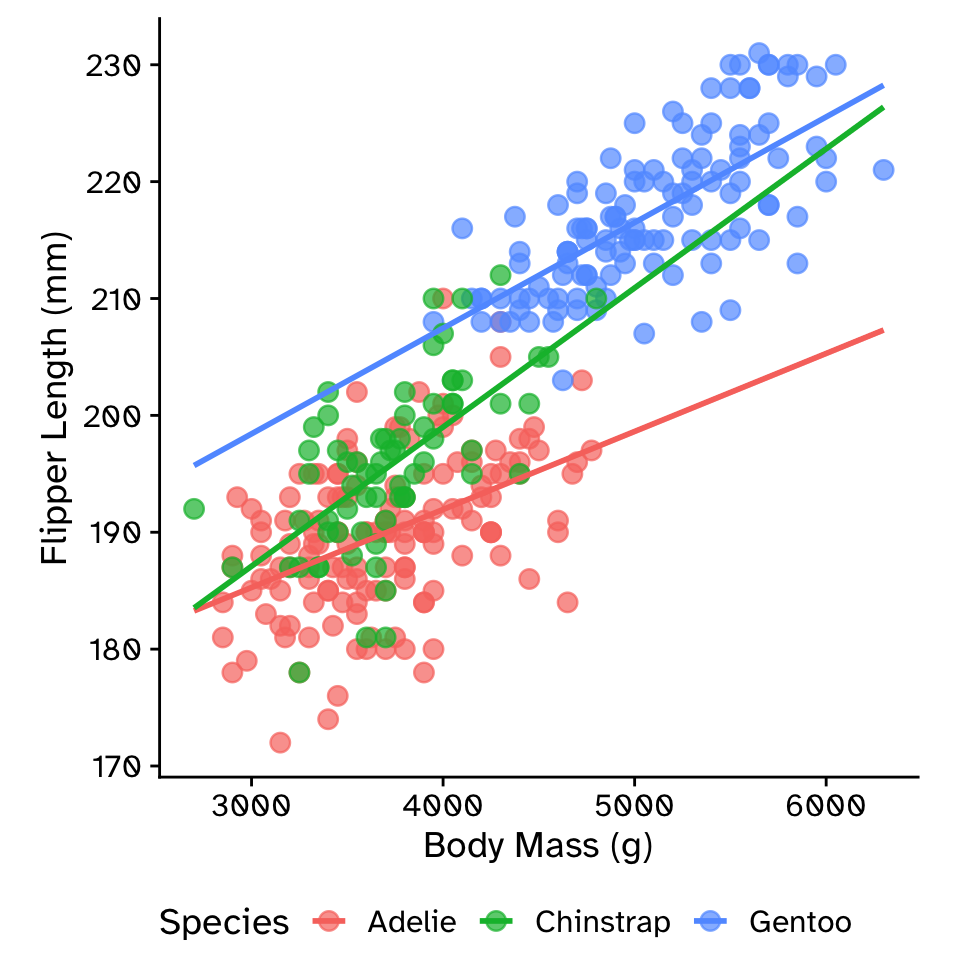
\[ \text{flipper_length_mm} = \beta_0 + \\ \beta_1 \times \text{body_mass_g} + \\ \beta_2 \times \text{species} + \\ \beta_3 \times \text{body_mass_g} \times \text{species} \]
Linear Regression
Multiple linear regression
- The methods used in
inferfor hypothesis testing in multiple linear regression are a bit limited.- Need to move to
parsnippackage or base Rlm()function. - Bit beyond this course, but you will learn it when you need to (hard to avoid!)
- Need to move to
- Super powerful framework
Logistic Regression
- When your response is a binomial (categorical variable with two levels), and you want to predict it using (at least) one quantitative variable
Logistic Regression
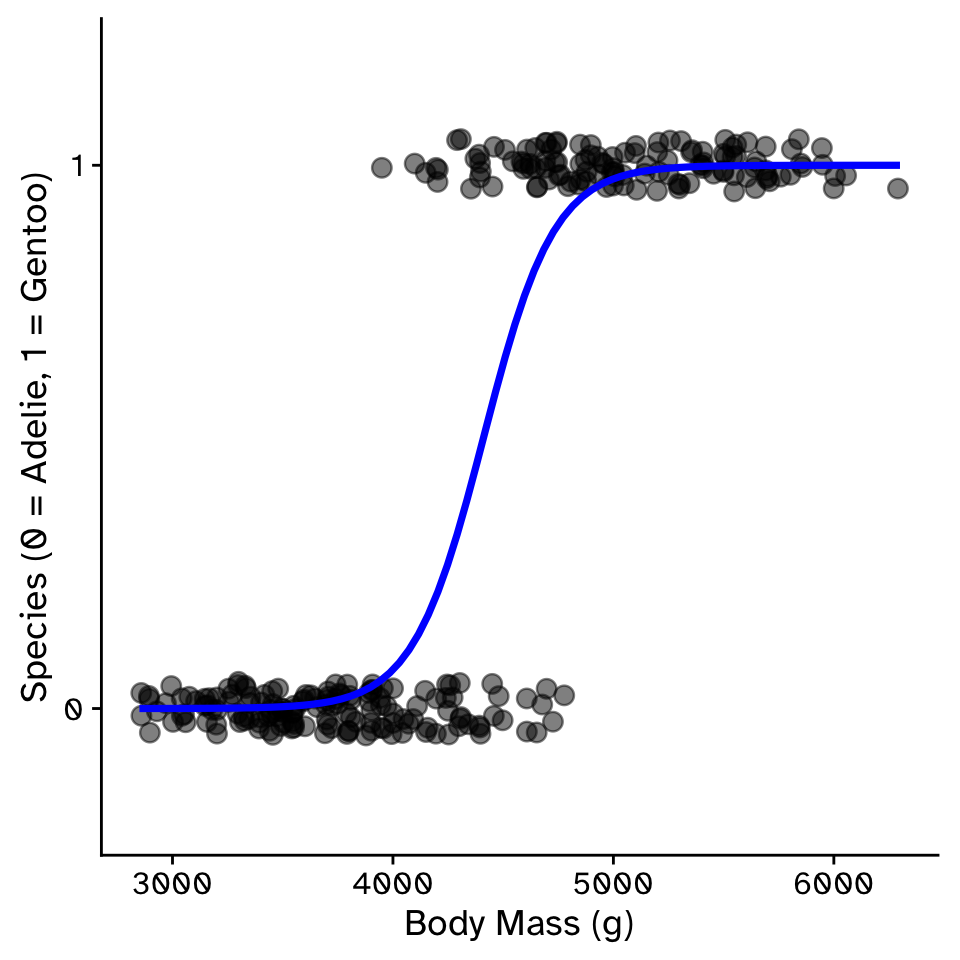
Predicting a binomial response with a quantitative variable
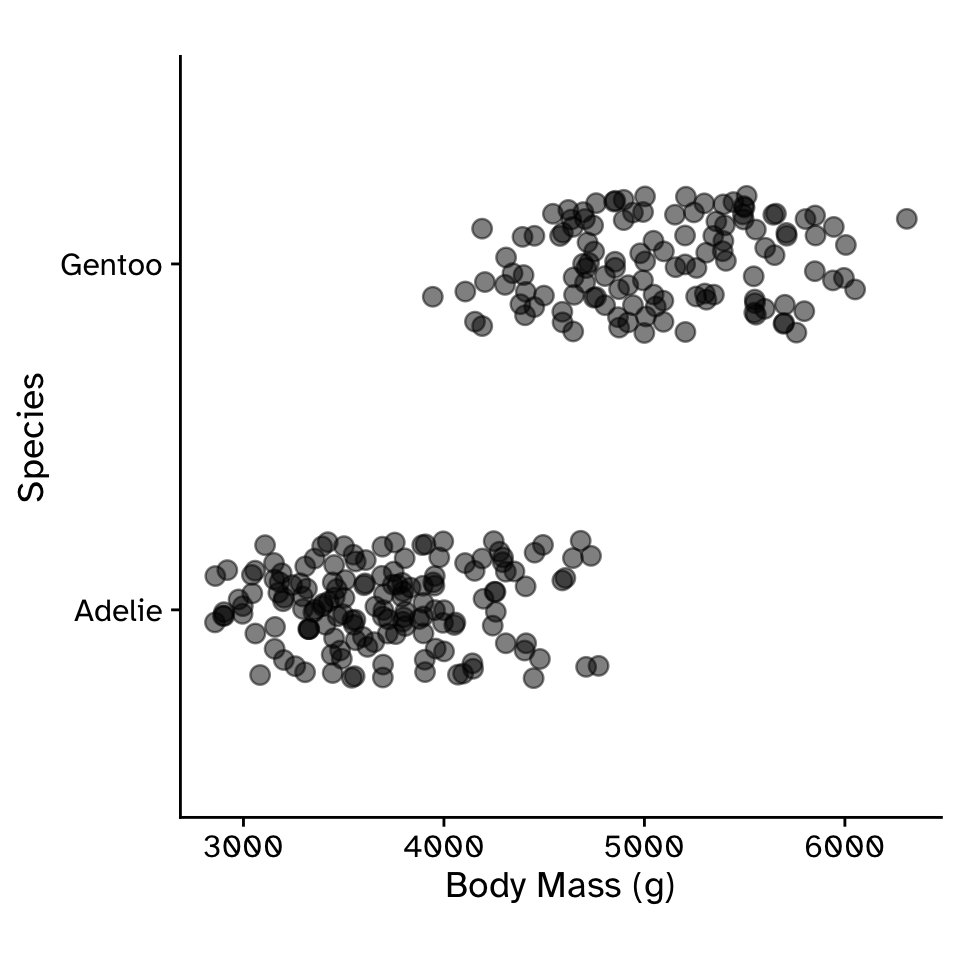
Logistic Regression
Predicting a binomial response with a quantitative variable
- Type of generalized linear model (GLM)
- Two categories are treated as 0 or 1.
- Requires a transformation of the response variable, the probability of being 1, \(p\)
- Linear regression is a straight line
- Will predict values less than 0 or more than 1
Logistic Regression
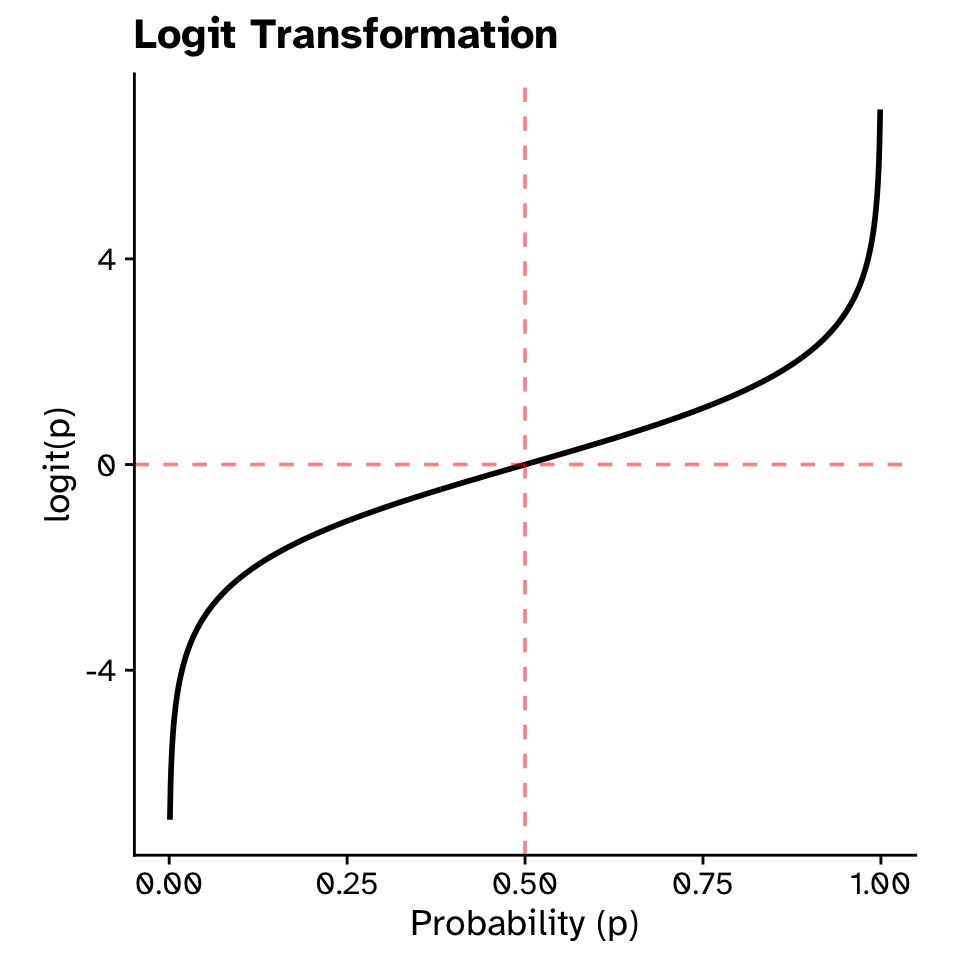
Logit transformation
\[ \text{logit}(p) = \log\left(\frac{p}{1-p}\right) \]
- When \(p=0.5\) (equally likely either species), \(\text{logit}(p)=0\)
- When \(p\) approaches 1 (almost certainly Gentoo), \(\text{logit}(p)\) approaches \(+\infty\)
- When \(p\) approaches 0 (almost certainly Adelie), \(\text{logit}(p)\) approaches \(-\infty\)
\[ \text{logit}(p) = \beta_0+\beta_1x+\beta_2x_2+... \]
Logistic Regression
Logit transformation
Logistic Regression
Inverse logit transformation
- After fitting a logistic regression model on the logit scale, you get predictions in logit units.
- The inverse logit transforms them back to probabilities (0–1).
\[ p = \frac{e^{\beta_0+\beta_1x...}}{1 + e^{\beta_0+\beta_1x...}} \]
Logistic Regression
How to fit in R
Logistic Regression
Interpreting coefficients: intercept
\(\beta_0 \approx -27.5\)
- Predicted logit when
body_mass_g = 0 - When body mass = 0 grams: \(p \approx 0\) (almost certainly Adelie)
- Rarely interpretable (0 grams is biologically impossible)
Logistic Regression
Interpreting coefficients: slope
\(\beta_1 \approx 0.00623\)
- For each additional gram of body mass, log-odds increase by 0.00623
- Odds ratio: \(e^{0.00623} \approx 1.0062\)
- Each gram of body mass multiplies the odds of being Gentoo by 1.0062
- A 100 gram difference: \(1.0062^{100} \approx 1.87\) times higher odds }
Logistic Regression
Interpreting coefficients
\[ \text{logit}(p) = -27.5 + 0.00623 \times \text{body_mass_g} \]
Inverse logit (to get probability): \[ p = \frac{e^{\text{logit}(p)}}{1 + e^{\text{logit}(p)}} \]
Logistic Regression
Multiple logisitic regression
- The methods used in
inferfor hypothesis testing in multiple logistic regression are a bit limited.- Need to move to
parsnippackage. - Bit beyond this course, but you will learn it when you need to (hard to avoid!)
- Need to move to
- Super powerful framework